Tested Applications
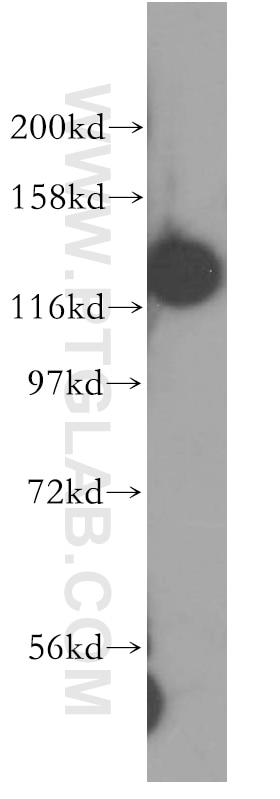
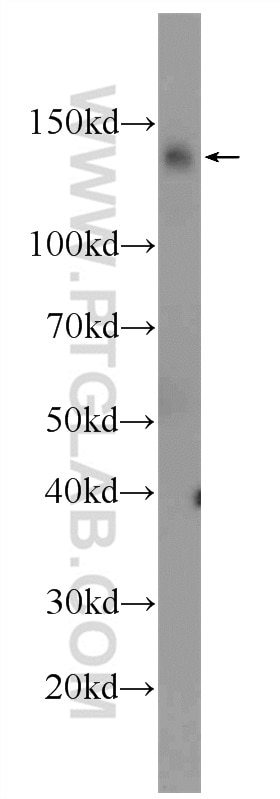
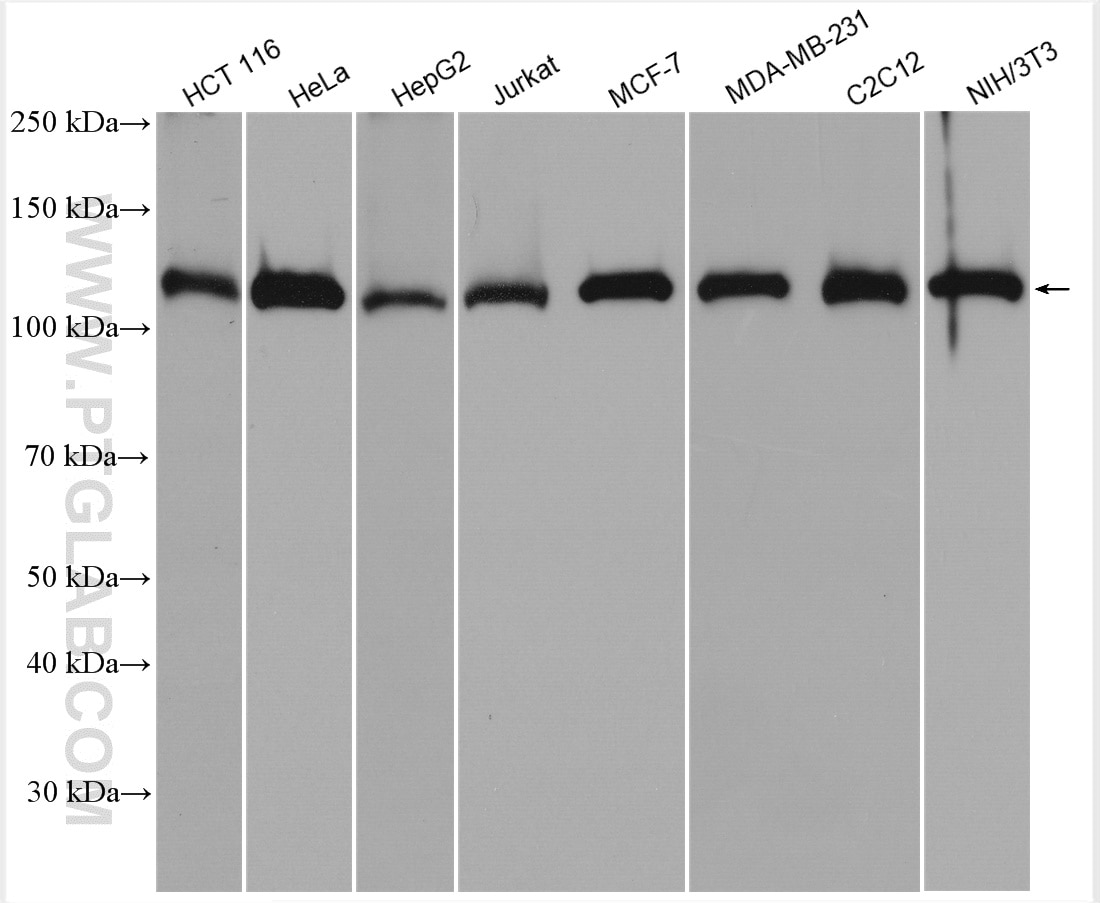
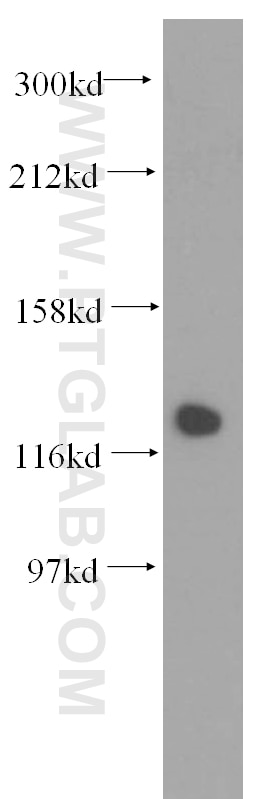
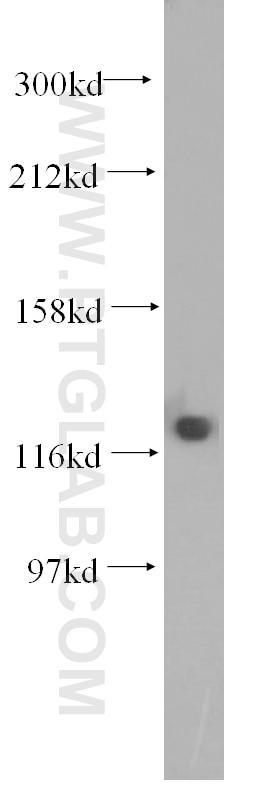
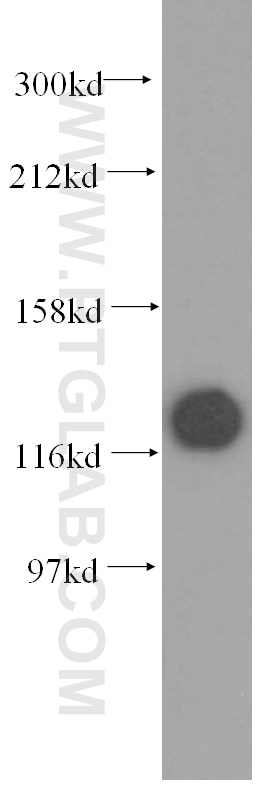
| Positive WB detected in | HCT 116 cells, mouse testis tissue, human kidney tissue, human placenta tissue, human brain tissue, rat testis tissue, HeLa cells, HepG2 cells, Jurkat cells, MCF-7 cells, MDA-MB-231 cells, C2C12 cells, NIH/3T3 cells |
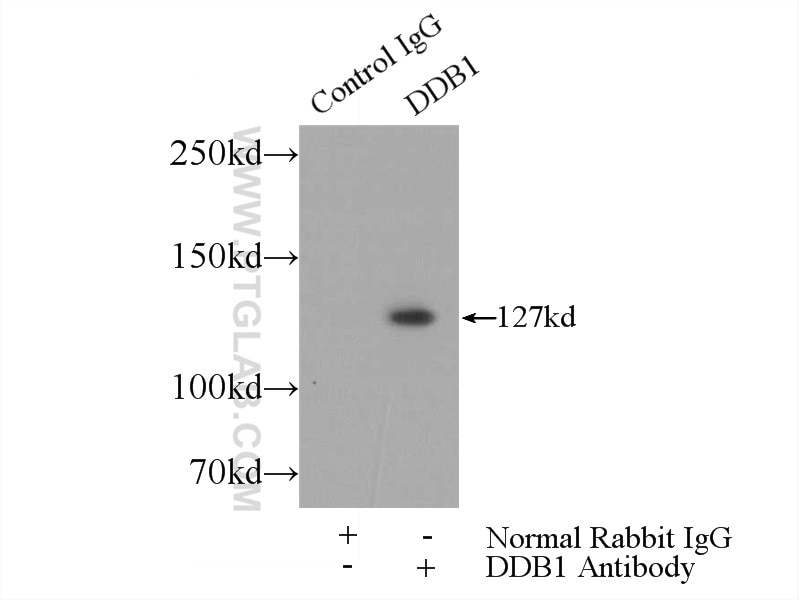
| Positive IP detected in | Jurkat cells |
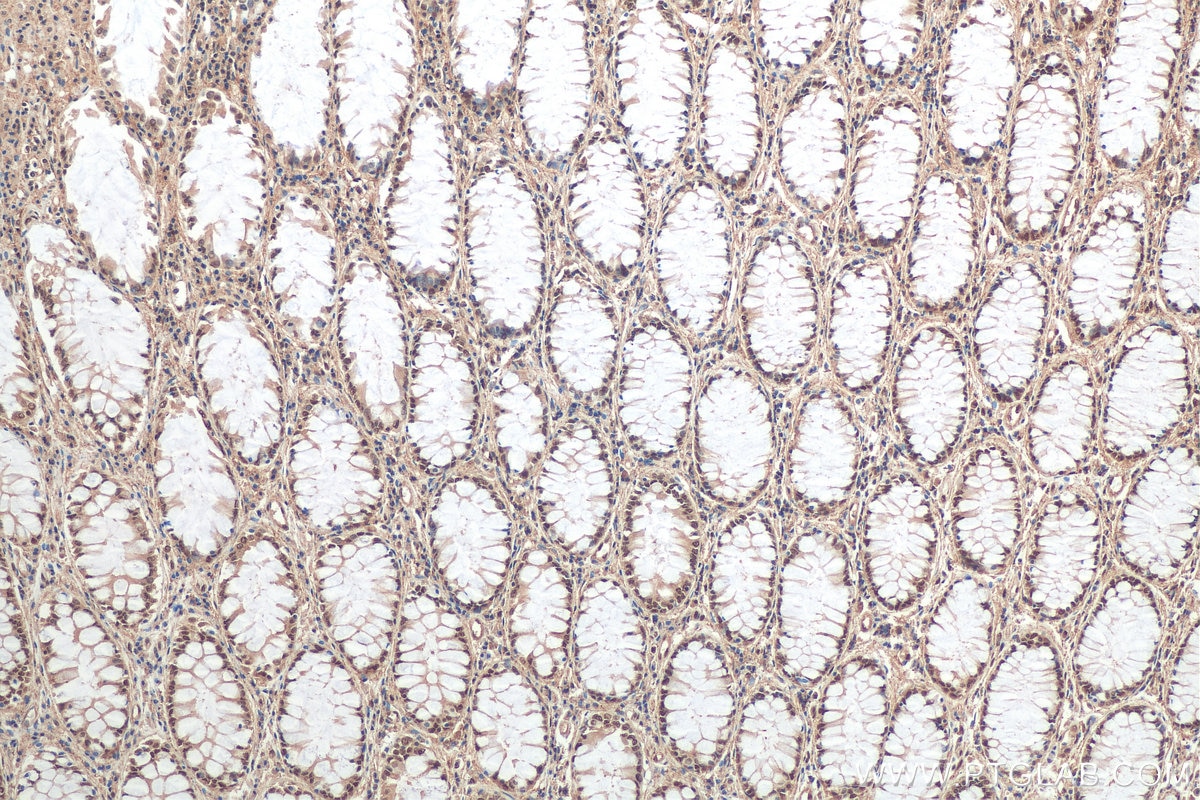
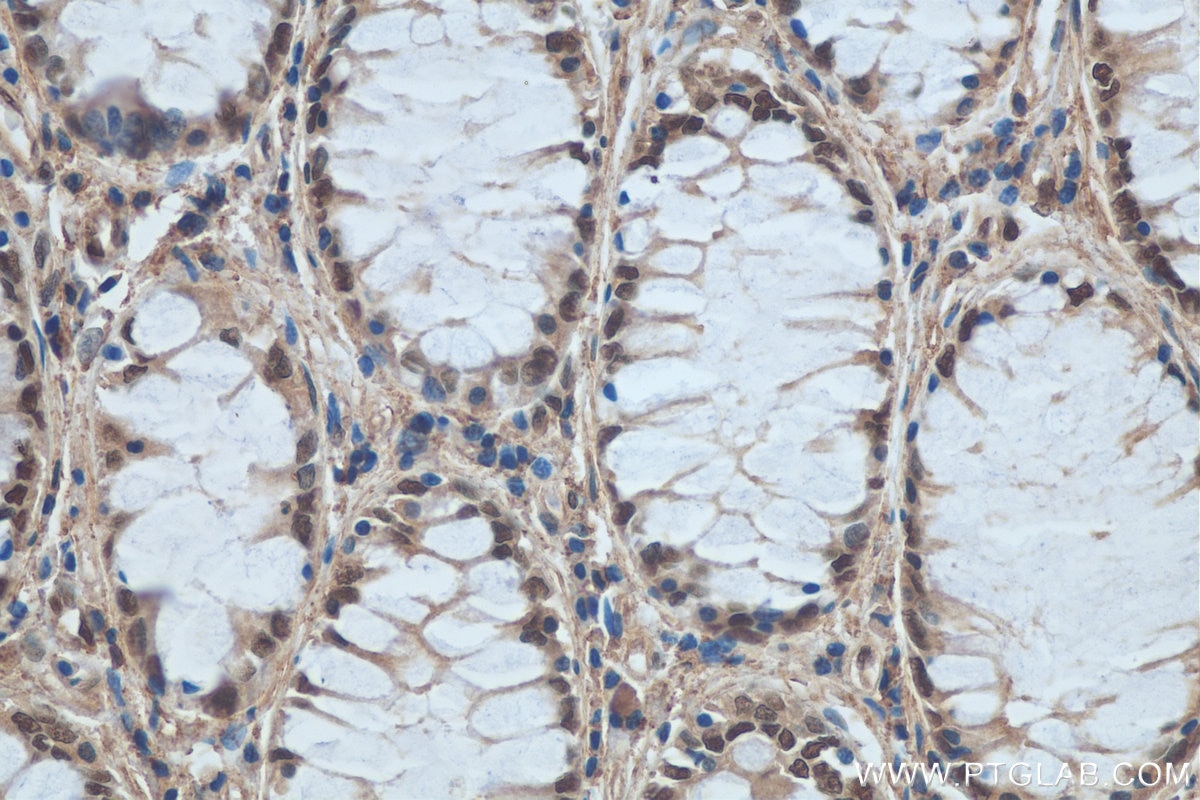
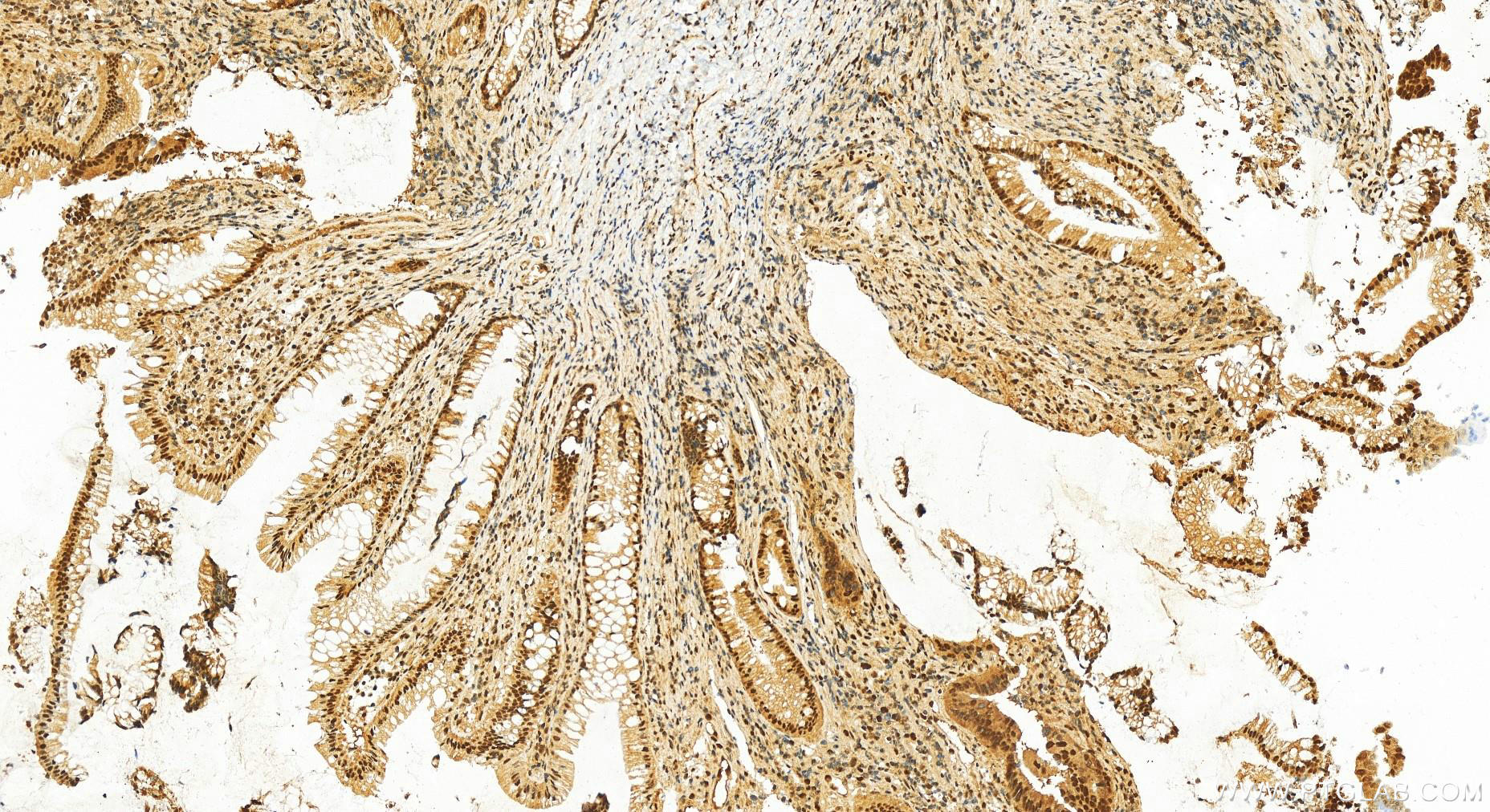
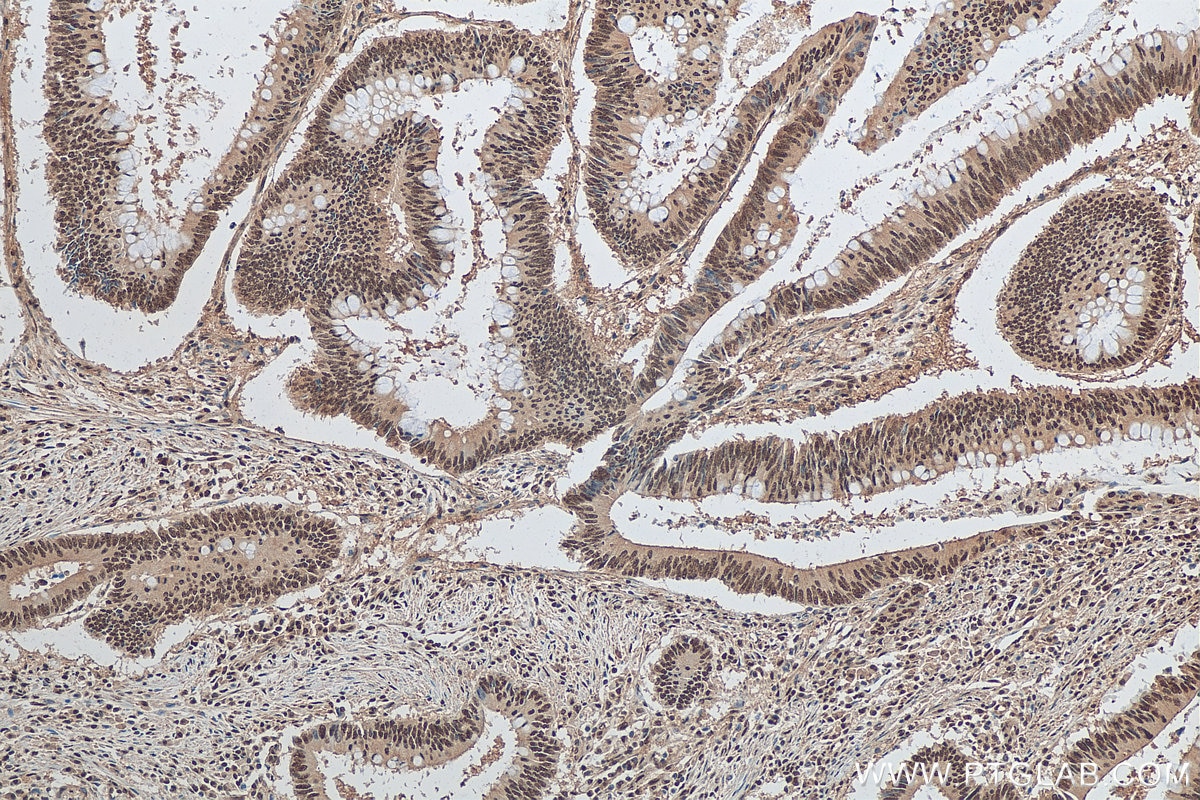
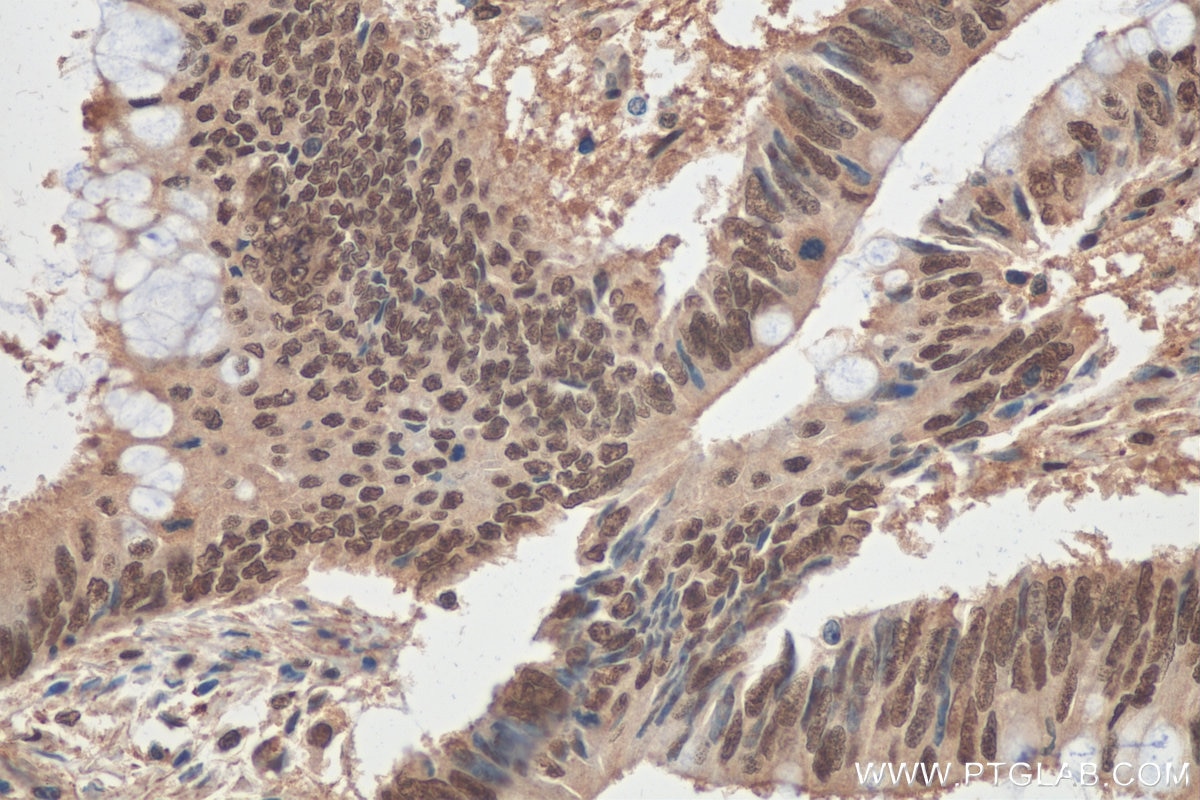
| Positive IHC detected in | human colon cancer tissue, human colon cancer Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:2000-1:16000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Immunohistochemistry (IHC) | IHC : 1:50-1:500 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 3 publications below |
| WB | See 14 publications below |
| IF | See 2 publications below |
| IP | See 1 publications below |
| CoIP | See 1 publications below |
| ChIP | See 1 publications below |
Product Information
11380-1-AP targets DDB1 in WB, IHC, IF, IP, CoIP, ChIP, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag1901 Product name: Recombinant human DDB1 protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 1-400 aa of BC011686 Sequence: MSYNYVVTAQKPTAVNGCVTGHFTSAEDLNLLIAKNTRLEIYVVTAEGLRPVKEVGMYGKIAVMELFRPKGESKDLLFILTAKYNACILEYKQSGESIDIITRAHGNVQDRIGRPSETGIIGIIDPECRMIGLRLYDGLFKVIPLDRDNKELKAFNIRLEELHVIDVKFLYGCQAPTICFVYQDPQGRHVKTYEVSLREKEFNKGPWKQENVEAEASMVIAVPEPFGGAIIIGQESITYHNGDKYLAIAPPIIKQSTIVCHNRVDPNGSRYLLGDMEGRLFMLLLEKEEQMDGTVTLKDLRVELLGETSIAECLTYLDNGVVFVGSRLGDSQLVKLNVDSNEQGSYVVAMETFTNLGPIVDMCVVDLERQGQGQLVTCSGAFKEGSLRIIRNGIGIHEHA Predict reactive species |
| Full Name | damage-specific DNA binding protein 1, 127kDa |
| Calculated Molecular Weight | 1140 aa, 127 kDa |
| Observed Molecular Weight | 127 kDa |
| GenBank Accession Number | BC011686 |
| Gene Symbol | DDB1 |
| Gene ID (NCBI) | 1642 |
| RRID | AB_2088808 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q16531 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
DDB1, also named as XAP1, XPCe, DDBa and XPE-BF, belongs to the DDB1 family. It is required for DNA repair. DDB1 binds to DDB2 to form the UV-damaged DNA-binding protein complex (the UV-DDB complex). The UV-DDB complex may recognize UV-induced DNA damage and recruit proteins of the nucleotide excision repair pathway (the NER pathway) to initiate DNA repair. The functional specificity of the DCX E3 ubiquitin-protein ligase complex is determined by the variable substrate recognition component recruited by DDB1. This antibody is specific to DDB1.
Protocols
| Product Specific Protocols | |
|---|---|
| IHC protocol for DDB1 antibody 11380-1-AP | Download protocol |
| IP protocol for DDB1 antibody 11380-1-AP | Download protocol |
| WB protocol for DDB1 antibody 11380-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Front Immunol Ddb1 Is Essential for the Expansion of CD4+ Helper T Cells by Regulating Cell Cycle Progression and Cell Death. | ||
FASEB J DDB2 promotes melanoma cell growth by transcriptionally regulating the expression of KMT2A and predicts a poor prognosis | ||
iScience A Multidimensional Characterization of E3 Ubiquitin Ligase and Substrate Interaction Network. | ||
PLoS One 14-3-3ε mediates the cell fate decision-making pathways in response of hepatocellular carcinoma to Bleomycin-induced DNA damage. | ||
Virology DDB1 is a cellular substrate of NS3/4A protease and required for hepatitis C virus replication.
| ||
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH Bernadette (Verified Customer) (09-24-2025) | The antibody worked well at 1:2000 dilution in 5% milk shaken overnight in 4 degree fridge on HEK293 cells.
|
FH Sarah (Verified Customer) (07-03-2019) | Total cell lysate (15 ug) was resolved on a 4-12% Bis-Tris gel and transferred to nitrocellulose membrane. Membrane was incubated in blocking buffer (5% milk/0.1% Tween-20) for 1h. Membrane was incubated with anti-DDB1 in blocking buffer (1:1000) at 4C overnight. After washing, membrane was incubated in anti-rabbit-HRP in blocking bufffer (1:3000) for 1h at room temperature. Protein was detected using ECL reagent and imaged on a chemiluminescence detection system.
 |