Tested Applications
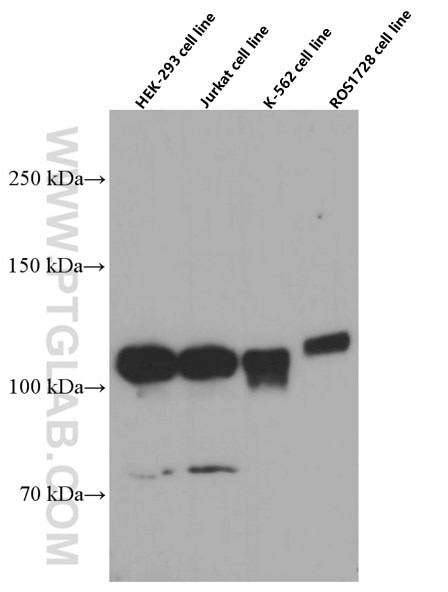
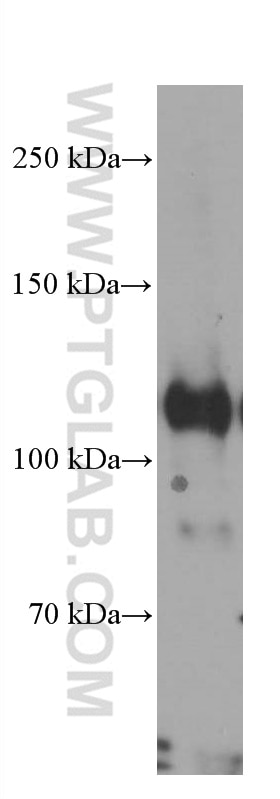
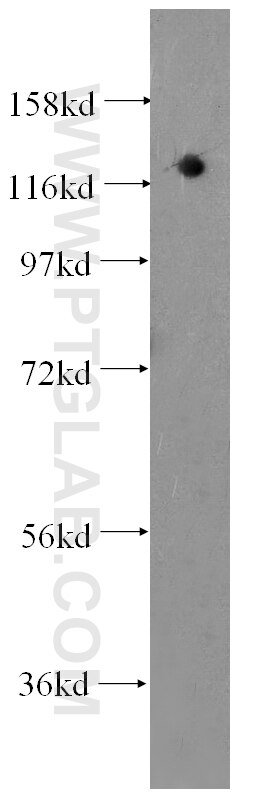
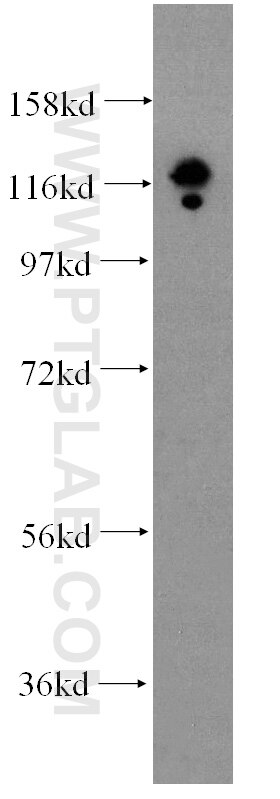
| Positive WB detected in | HEK-293 cells, HepG2 cells, BxPC-3 cells, RAW 264.7 cells, A431 cells, Jurkat cells, K-562 cells, ROS1728 cells |
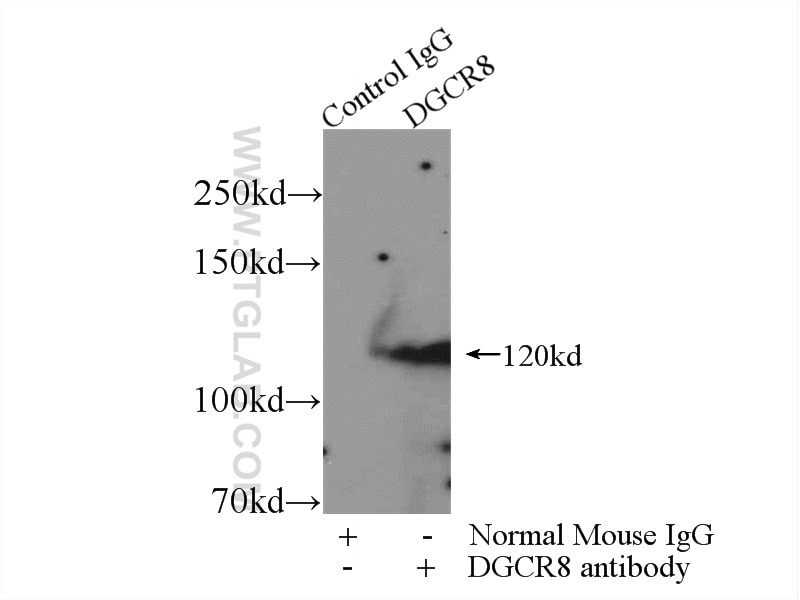
| Positive IP detected in | K-562 cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:1000-1:6000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 1 publications below |
| WB | See 13 publications below |
| IP | See 1 publications below |
| RIP | See 2 publications below |
Product Information
60084-1-Ig targets DGCR8 in WB, IP, RIP, ELISA applications and shows reactivity with human, mouse, rat, pig samples.
| Tested Reactivity | human, mouse, rat, pig |
| Cited Reactivity | human, mouse, rat, chicken |
| Host / Isotype | Mouse / IgG2a |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag4871 Product name: Recombinant human DGCR8 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 167-517 aa of BC009323 Sequence: VKAKVEVCKDESVDLEEFRSYLEKRFDFEQVTVKKFRTWAERRQFNREMKRKQAESERPILPANQKLITLSVQDAPTKKEFVINPNGKSEVCILHEYMQRVLKVRPVYNFFECENPSEPFGASVTIDGVTYGSGTASSKKLAKNKAARATLEILIPDFVKQTSEEKPKDSEELEYFNHISIEDSRVYELTSKAGLLSPYQILHECLKRNHGMGDTSIKFEVVPGKNQKSEYVMACGKHTVRGWCKNKRVGKQLASQKILQLLHPHVKNWGSLLRMYGRESSKMVKQETSDKSVIELQQYAKKNKPNLHILSKLQEEMKRLAEEREETRKKPKMSIVASAQPGGEPLCTVDV Predict reactive species |
| Full Name | DiGeorge syndrome critical region gene 8 |
| Calculated Molecular Weight | 773 aa, 86 kDa |
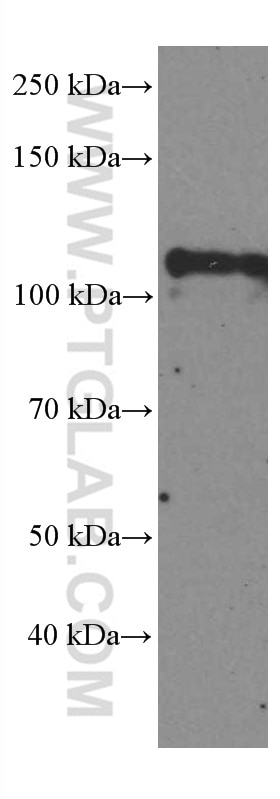
| Observed Molecular Weight | 120 kDa |
| GenBank Accession Number | BC009323 |
| Gene Symbol | DGCR8 |
| Gene ID (NCBI) | 54487 |
| RRID | AB_2090985 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein A purification |
| UNIPROT ID | Q8WYQ5 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
DGCR8 is a RNA-binding protein that assists the Rnase III enzyme Drosha in the processing of microRNAs (miRNAs), which regulate the expression of a large number of protein-coding genes[PMID: 22580560]. DGCR8, which contains two double-stranded RNA (dsRNA)-binding domains, may be an essential component of the primary miRNAs processing complex, along with Drosha, promoting the processing of primary microRNA to precursor microRNA. It is ubiquitous expressed in human and mouse tissues, and is deleted in DiGeorge syndrome[22323604]. The calculated molecular weight of DGCR8 is 82-86 kDa, but the post-modified DGCR8 is about 120 kDa.
Protocols
| Product Specific Protocols | |
|---|---|
| IP protocol for DGCR8 antibody 60084-1-Ig | Download protocol |
| WB protocol for DGCR8 antibody 60084-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Mol Cell A transient transcriptional activation governs unpolarized-to-polarized morphogenesis during embryo implantation | ||
Nat Commun Hypoxia regulates overall mRNA homeostasis by inducing Met1-linked linear ubiquitination of AGO2 in cancer cells. | ||
Aging Cell p38 MAPK-mediated loss of nuclear RNase III enzyme Drosha underlies amyloid beta-induced neuronal stress in Alzheimer's disease | ||
Mol Ther DGCR8/ZFAT-AS1 Promotes CDX2 Transcription in a PRC2 Complex-Dependent Manner to Facilitate the Malignant Biological Behavior of Glioma Cells. | ||
Nucleic Acids Res SUMOylation at K707 of DGCR8 controls direct function of primary microRNA. |