Tested Applications
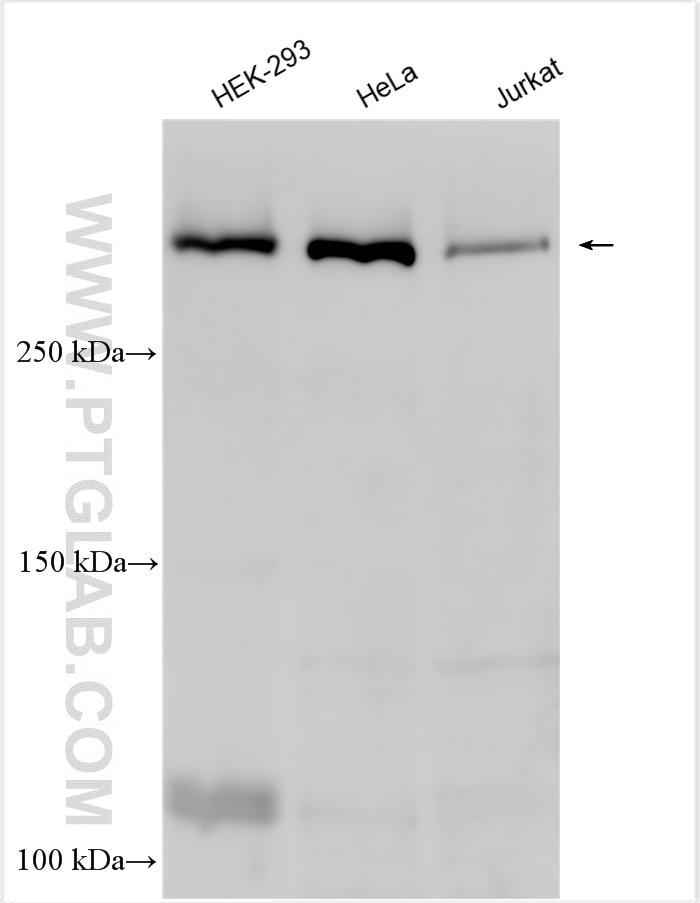
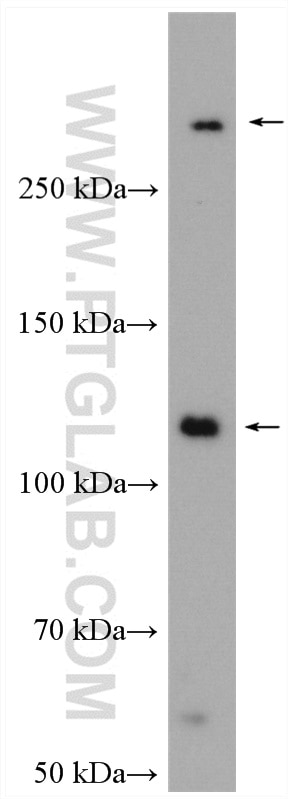
| Positive WB detected in | HEK-293 cells, A2780 cells, HeLa cells, HepG2 cells, Jurkat cells, OS-RC-2 cells |
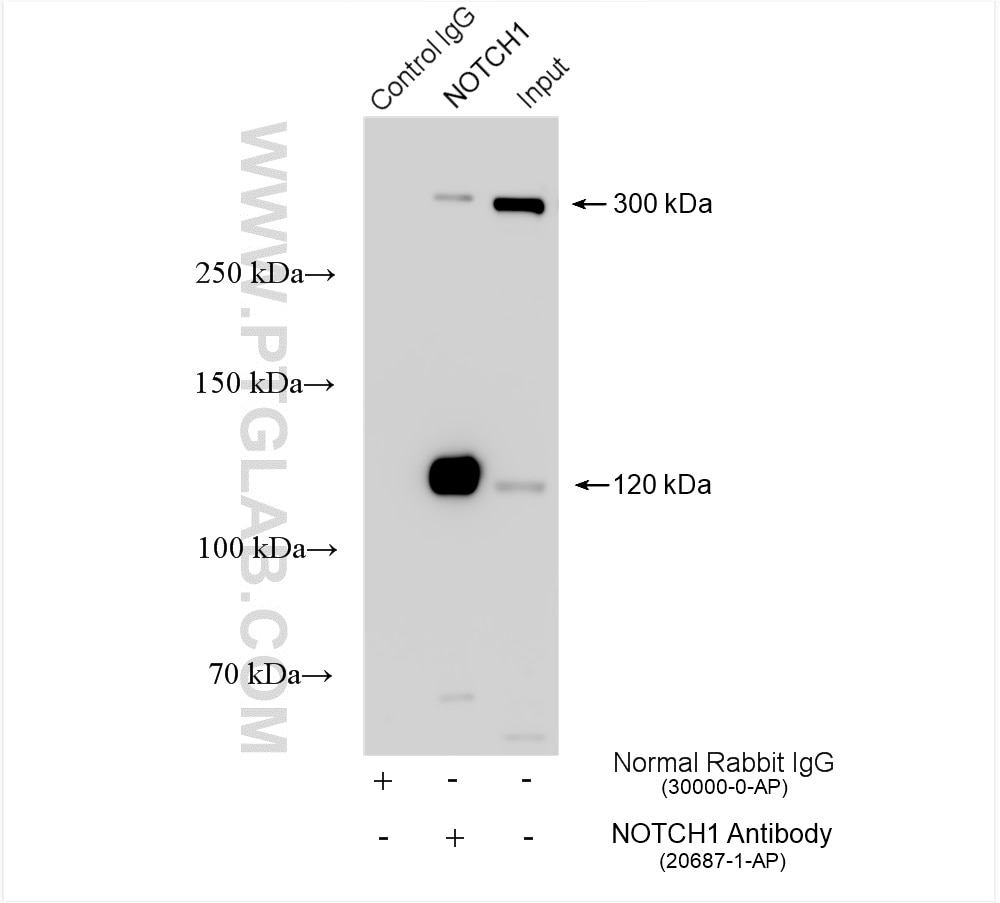
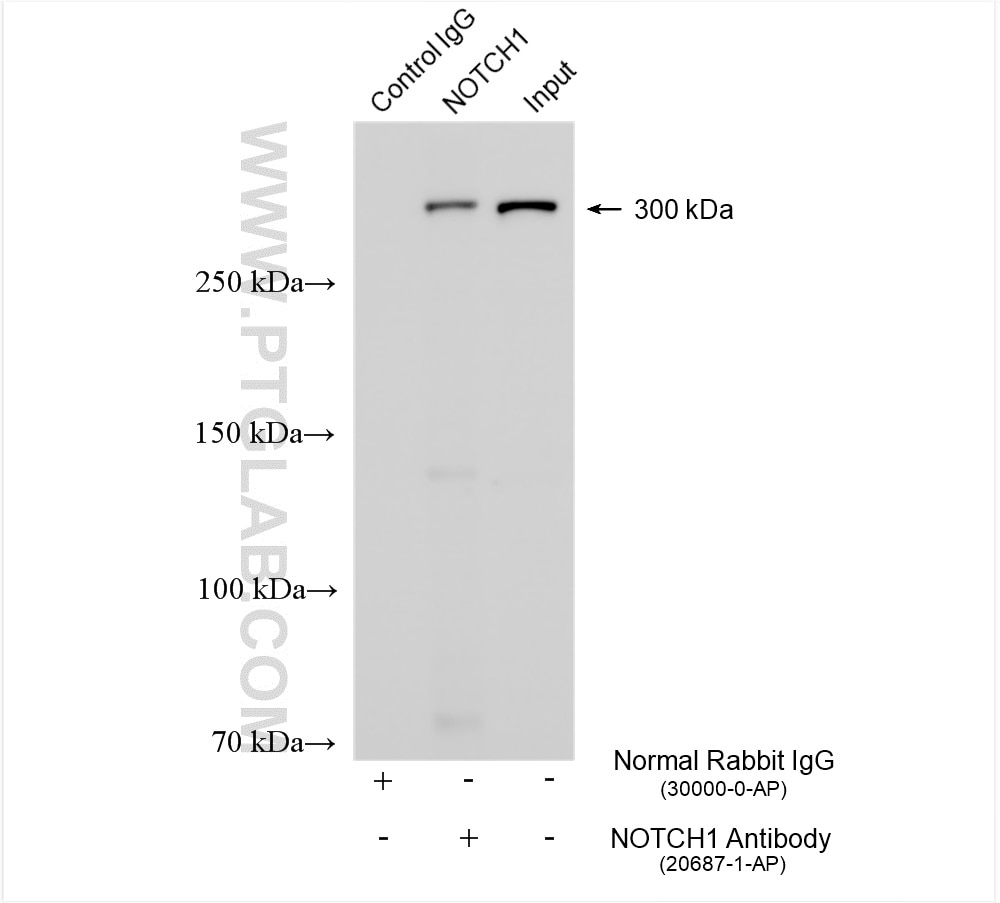
| Positive IP detected in | HEK-293 cells, HepG2 cells |
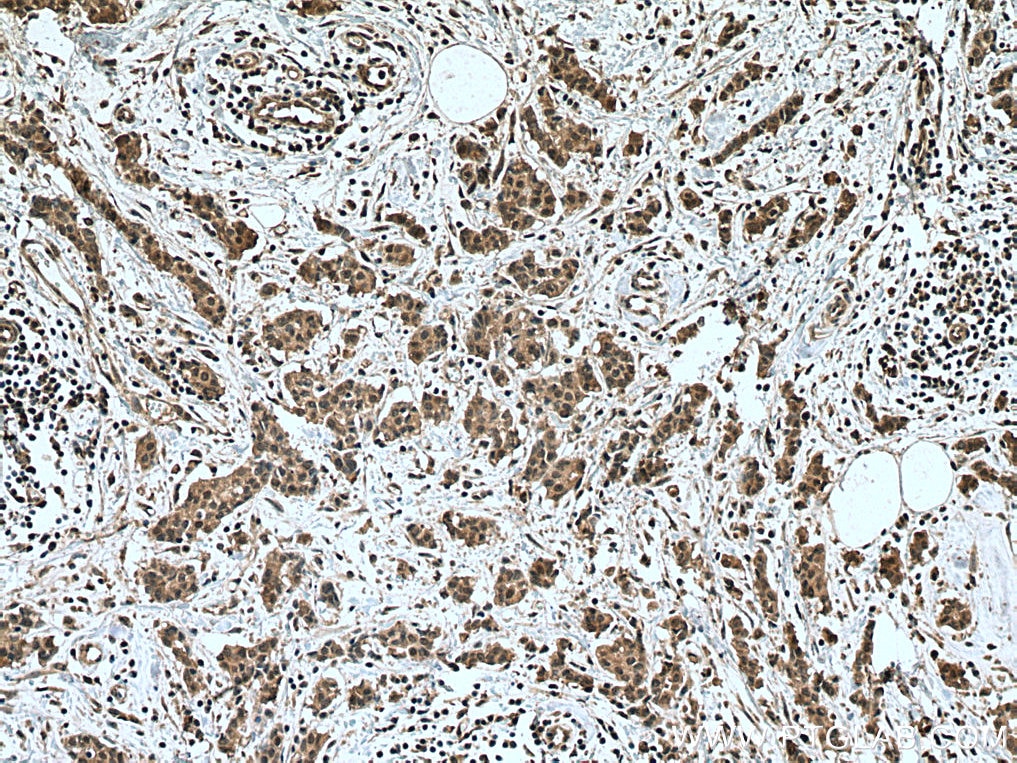
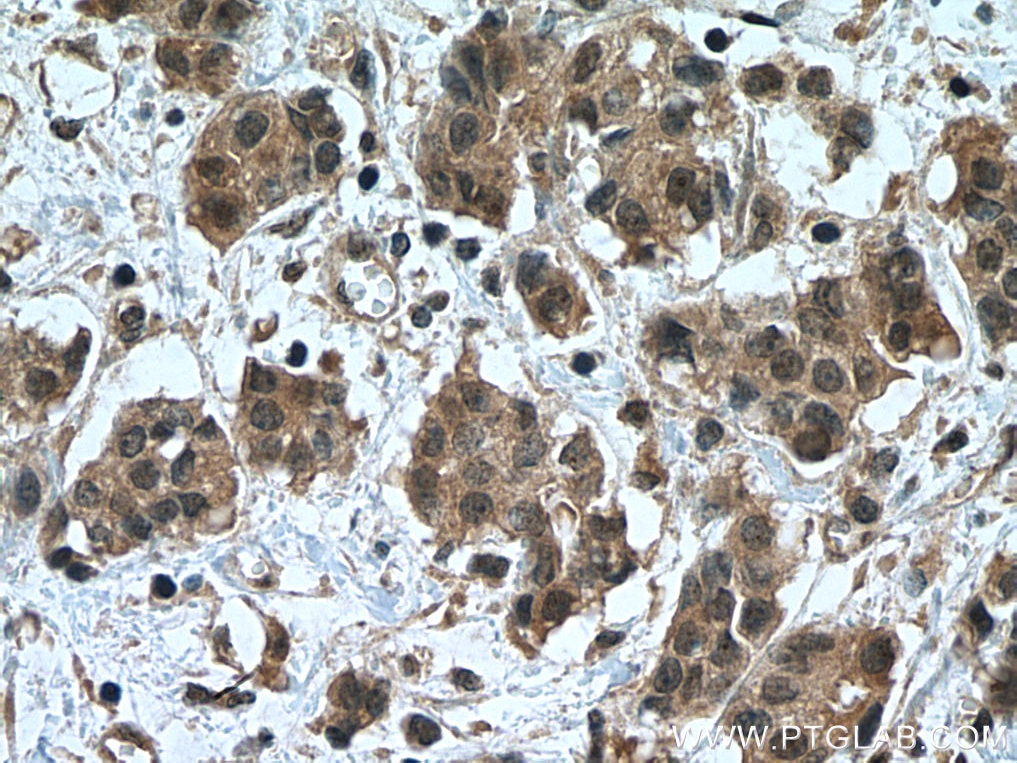
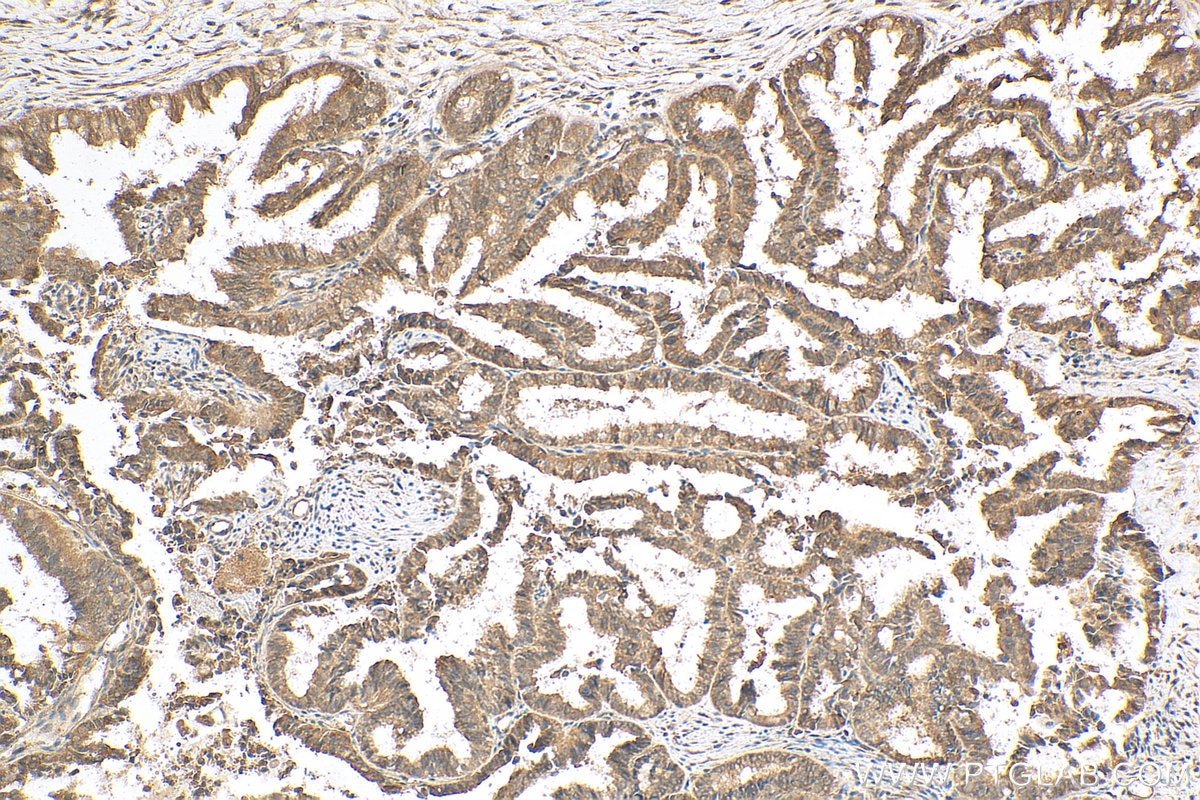
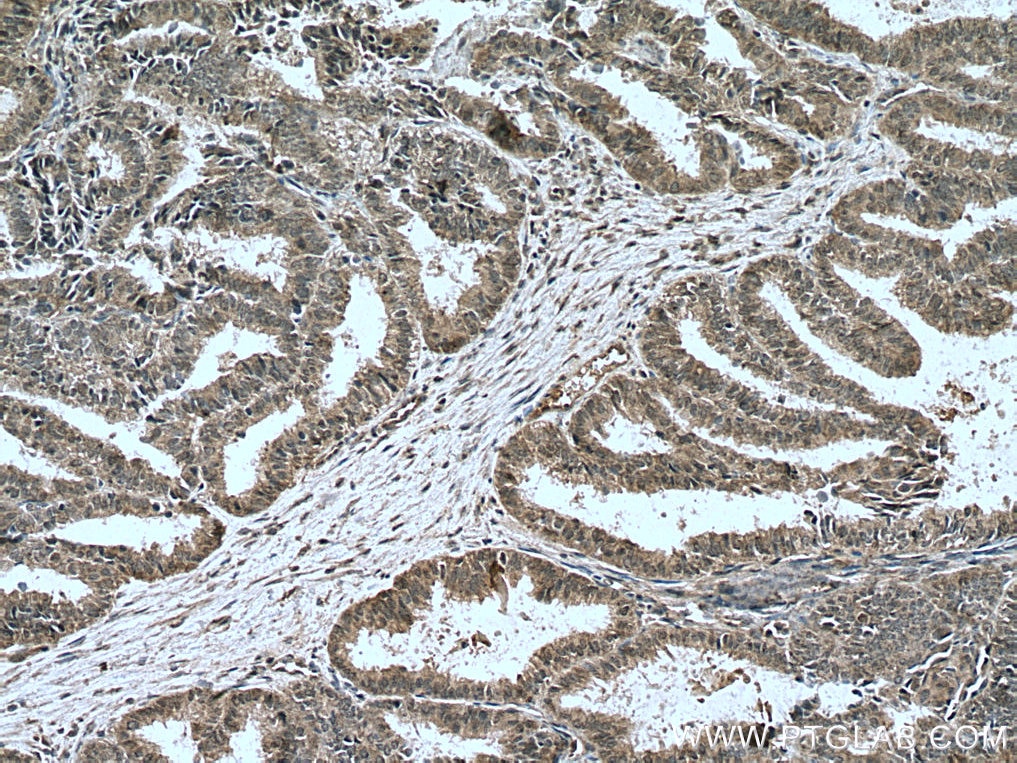
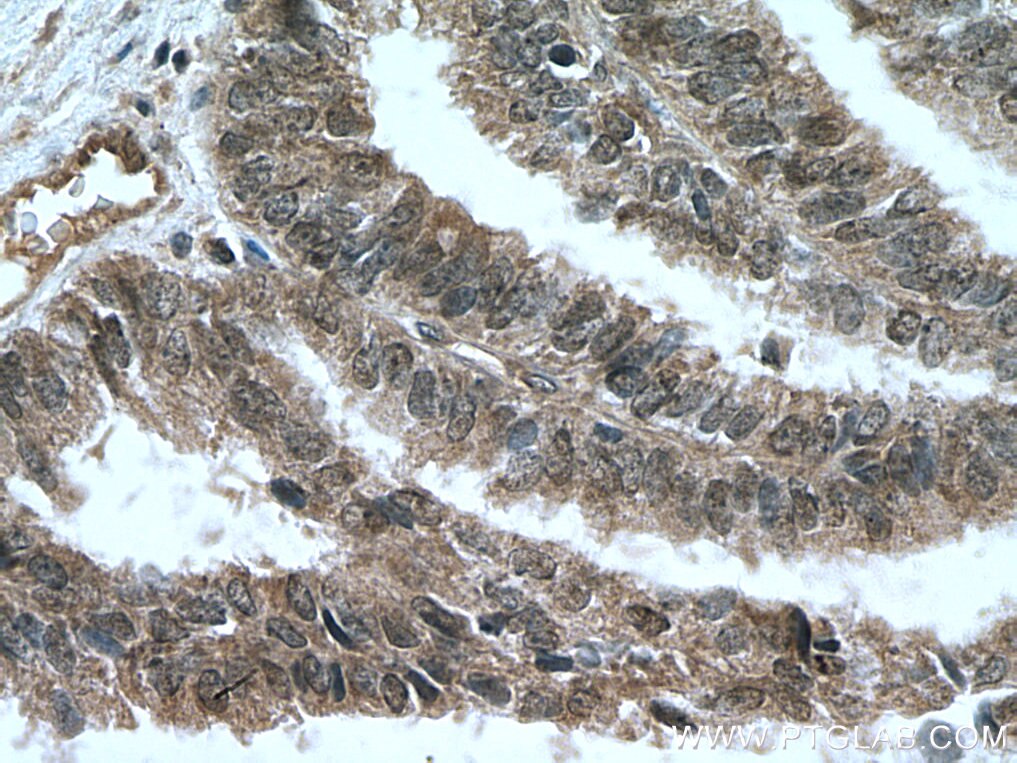
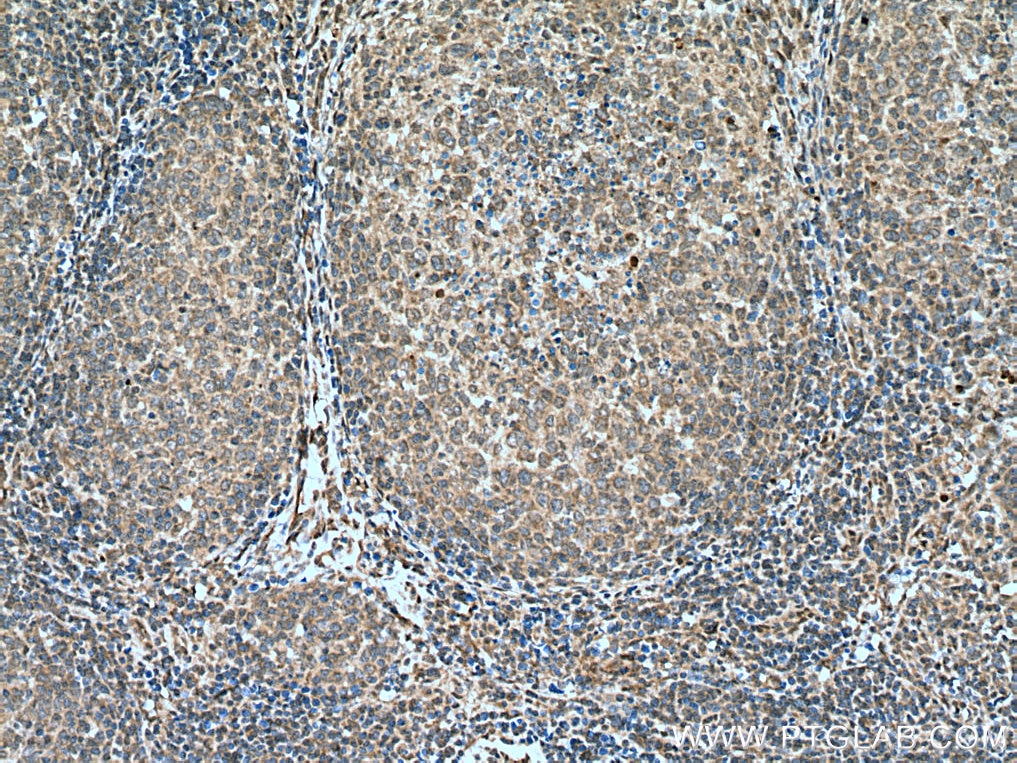
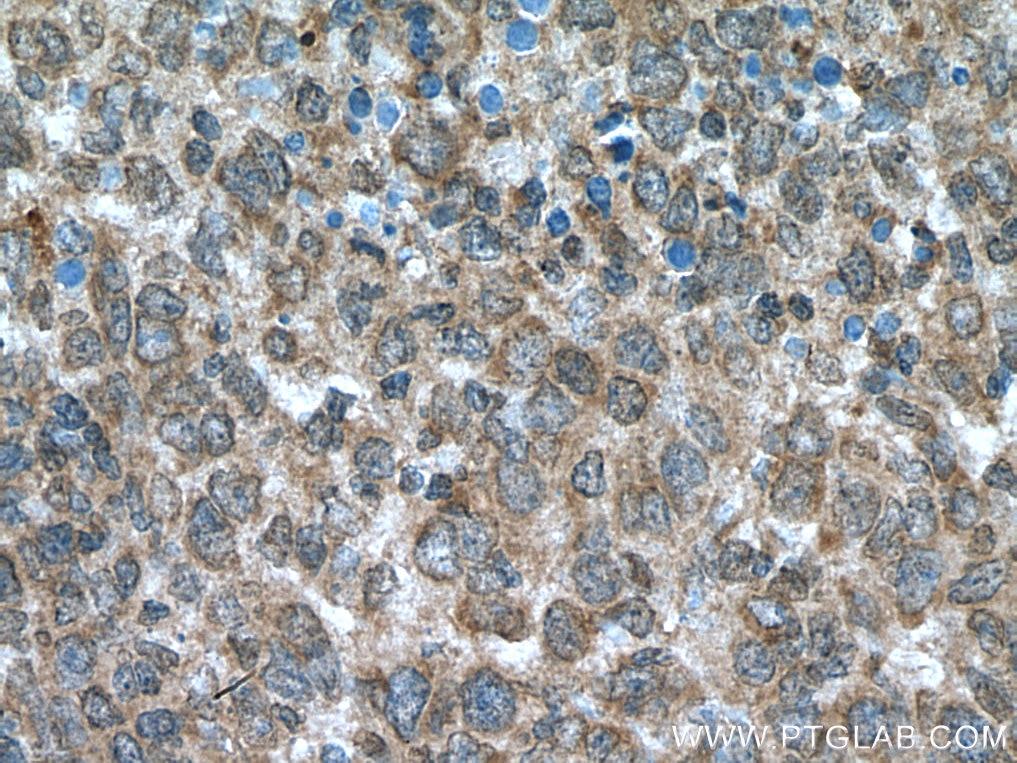
| Positive IHC detected in | human breast cancer tissue, human ovary tumor tissue, human lymphoma tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
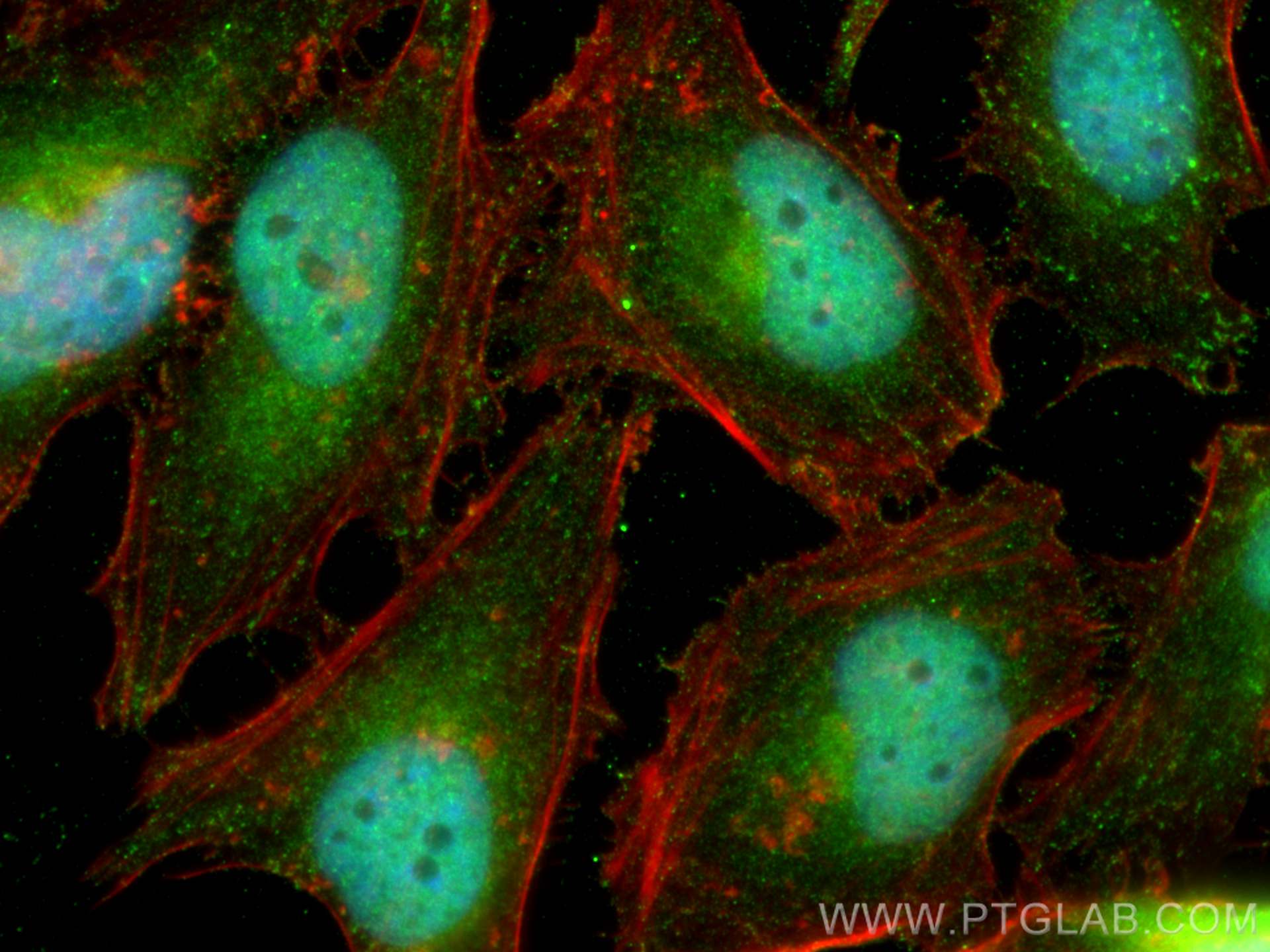
| Positive IF/ICC detected in | HeLa cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:500-1:1000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Immunohistochemistry (IHC) | IHC : 1:50-1:500 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:200-1:800 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 3 publications below |
| WB | See 102 publications below |
| IHC | See 23 publications below |
| IF | See 19 publications below |
| IP | See 1 publications below |
| CoIP | See 6 publications below |
Product Information
20687-1-AP targets NOTCH1 in WB, IHC, IF/ICC, IP, CoIP, ELISA applications and shows reactivity with human samples.
| Tested Reactivity | human |
| Cited Reactivity | human, pig, zebrafish, mink |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
Peptide Predict reactive species |
| Full Name | Notch homolog 1, translocation-associated (Drosophila) |
| Calculated Molecular Weight | 273 kDa |
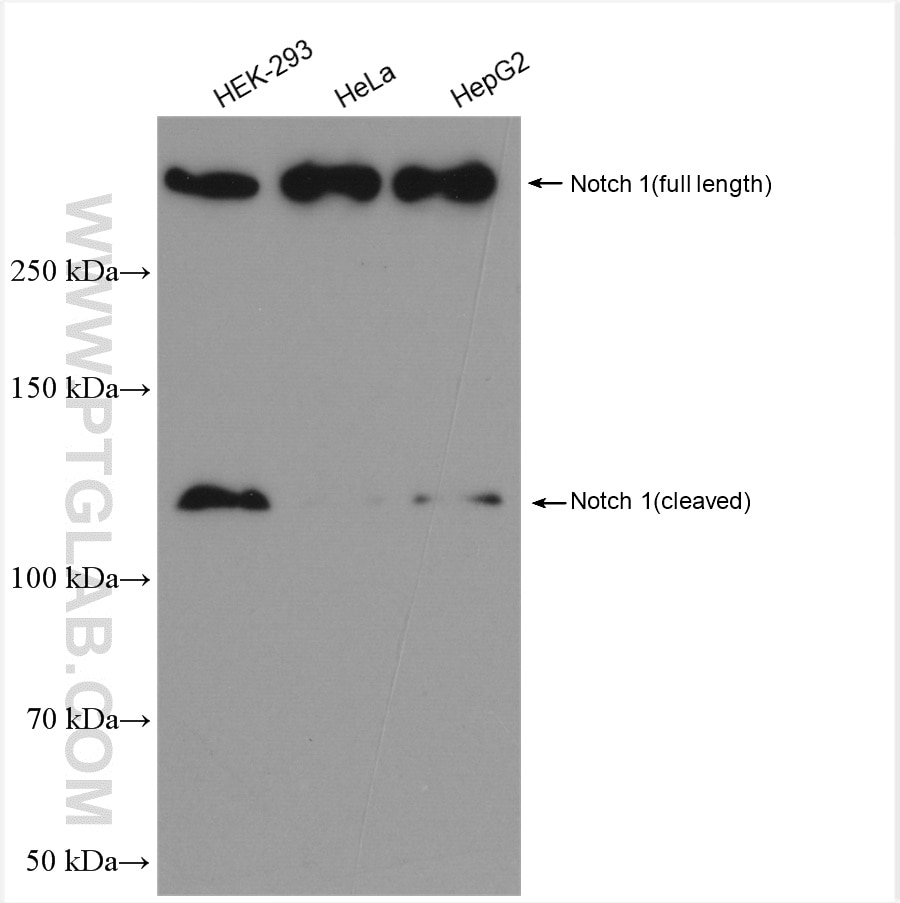
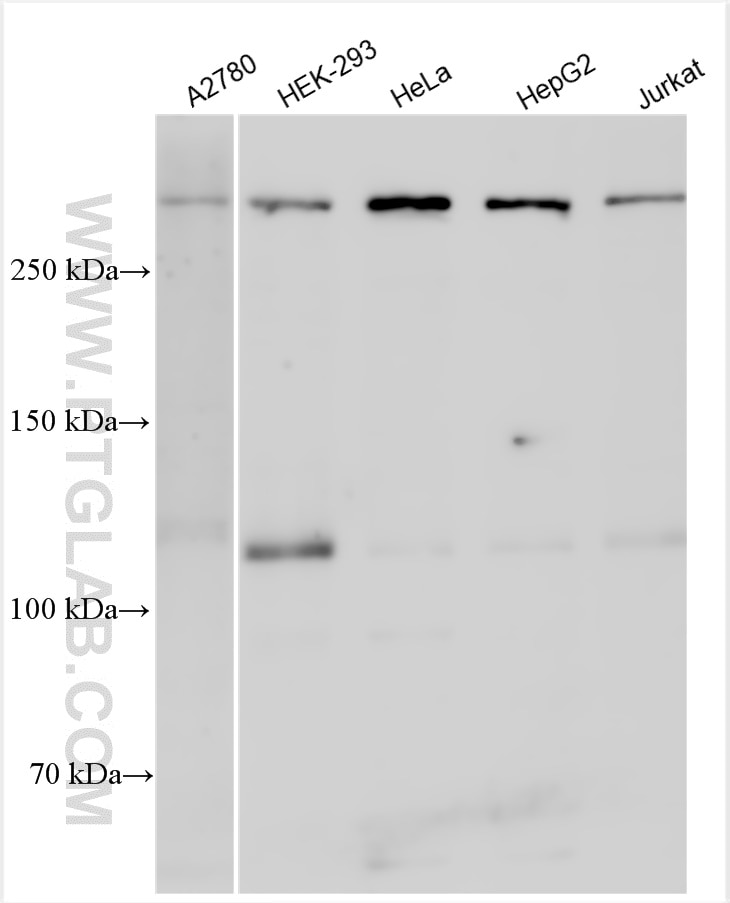
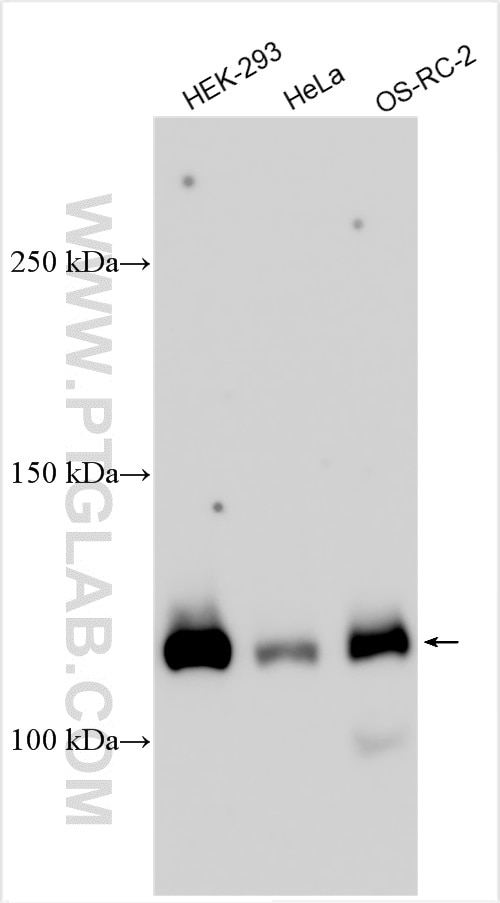
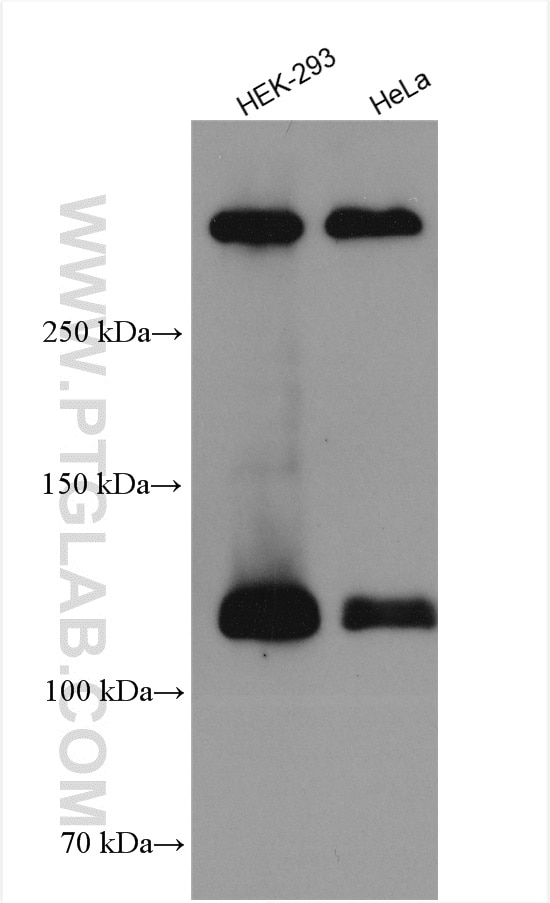
| Observed Molecular Weight | 273-300 kDa, 120 kDa |
| GenBank Accession Number | NM_017617 |
| Gene Symbol | NOTCH1 |
| Gene ID (NCBI) | 4851 |
| RRID | AB_10700012 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | P46531 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
NOTCH1, also named as TAN1, belongs to the NOTCH family. NOTCH1 functions as a receptor for membrane-bound ligands Jagged1, Jagged2 and Delta1 to regulate cell-fate determination. Upon ligand activation through the released notch intracellular domain (NICD) it forms a transcriptional activator complex with RBP-J kappa and activates genes of the enhancer of split locus. NOTCH1 affects the implementation of differentiation, proliferation and apoptotic programs. It may be important for normal lymphocyte function. In altered form, may contribute to transformation or progression in some T-cell neoplasms. NOTCH1 is involved in the maturation of both CD4+ and CD8+ cells in the thymus. May be important for follicular differentiation and possibly cell fate selection within the follicle. During cerebellar development, may function as a receptor for neuronal DNER and may be involved in the differentiation of Bergmann glia. Defects in NOTCH1 are a cause of bicuspid aortic valve (BAV).
Notch is synthesized in the endoplasmic reticulum as an inactive form which is proteolytically cleaved by a furin-like convertase (S1 cleavage) in the trans-golgi network before it reaches the plasma membrane to yield an active, ligand-accessible form. Cleavage results in a C-terminal fragment N(TM) and a N-terminal fragment N(EC). Following ligand binding, it is cleaved (S2 cleavage) by TNF-alpha converting enzyme (TACE) to yield a membrane-associated intermediate fragment called Notch extracellular truncation (NEXT). This fragment is then cleaved by presenilin-dependent gamma-secretase (S3 cleavage) to release the intracellular domain (NICD) from the membrane. The antibody is specific to NOTCH1. It can recognize the full length NOTCH1(270 kDa) and cleaved NOTCH1 form (120 kDa).
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for NOTCH1 antibody 20687-1-AP | Download protocol |
| IHC protocol for NOTCH1 antibody 20687-1-AP | Download protocol |
| IP protocol for NOTCH1 antibody 20687-1-AP | Download protocol |
| WB protocol for NOTCH1 antibody 20687-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cell Metab Mitochondrial Dynamics Is Critical for the Full Pluripotency and Embryonic Developmental Potential of Pluripotent Stem Cells. | ||
Mol Psychiatry Medial prefrontal cortex Notch1 signalling mediates methamphetamine-induced psychosis via Hes1-dependent suppression of GABAB1 receptor expression. | ||
EBioMedicine Immunological profiles of human oligodendrogliomas define two distinct molecular subtypes | ||
ACS Appl Mater Interfaces Advanced Strategies in Bone Tissue Engineering: "Membrane-Jelly" Hydrogel System to Improve Bone Marrow Stem Cell Osteogenic Differentiation and Bone Regeneration | ||
Cell Rep Transcriptional control of brain tumor stem cells by a carbohydrate binding protein |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH Lili (Verified Customer) (02-05-2026) | Great antibody. Works amazing for western blot. You can detect both NOTCH and NCID.
 |
FH Nikolett (Verified Customer) (10-14-2025) | Antibody is work on human lung cancer cell lines after overnight incubation, but the bands were a bit faint.
|