Tested Applications
| Positive WB detected in | HeLa cells, Jurkat cells, HEK-293 cells |
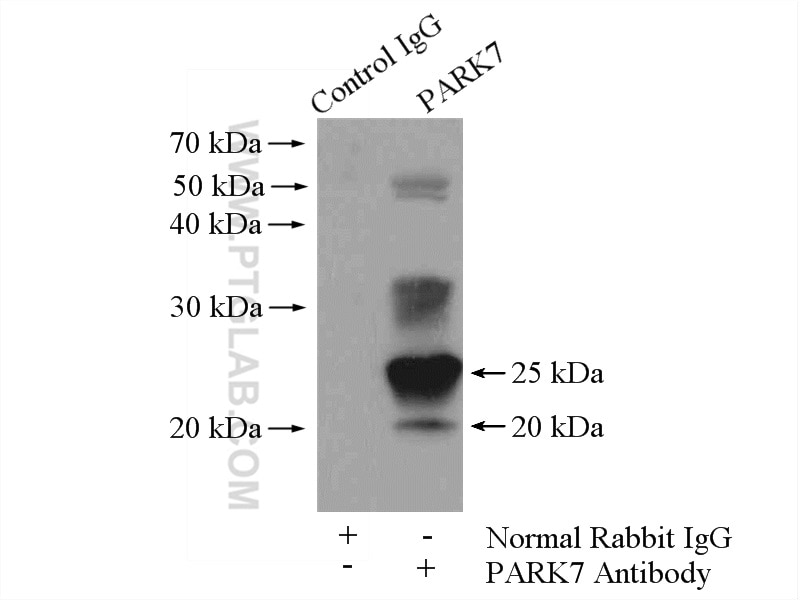
| Positive IP detected in | HeLa cells |
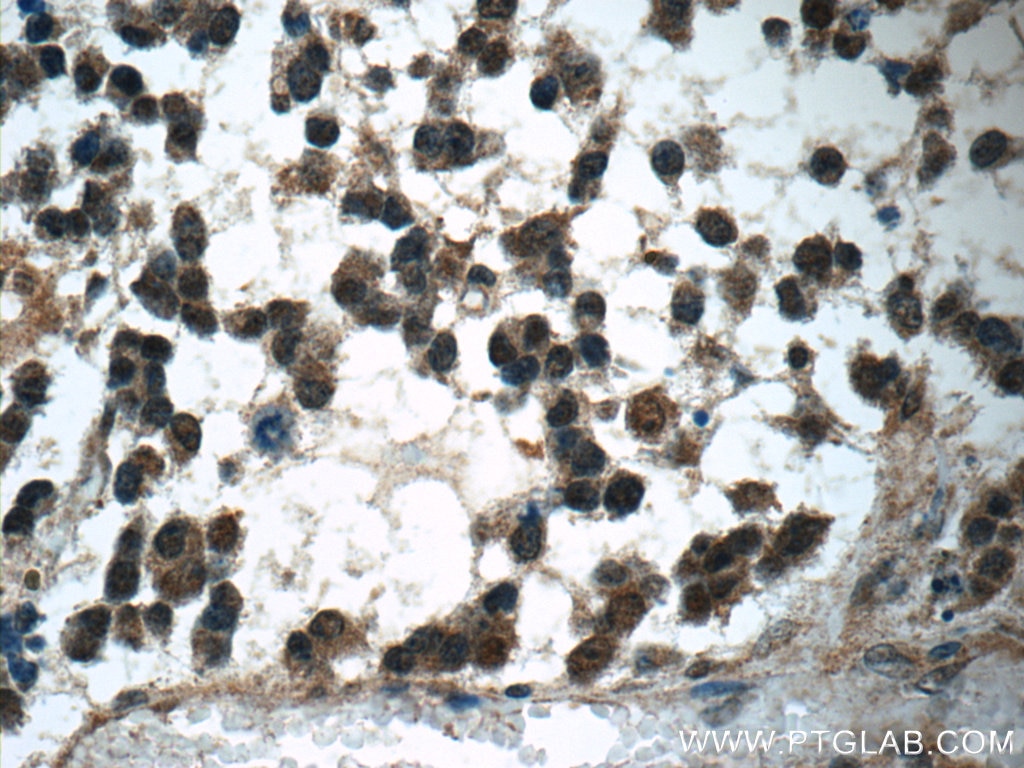
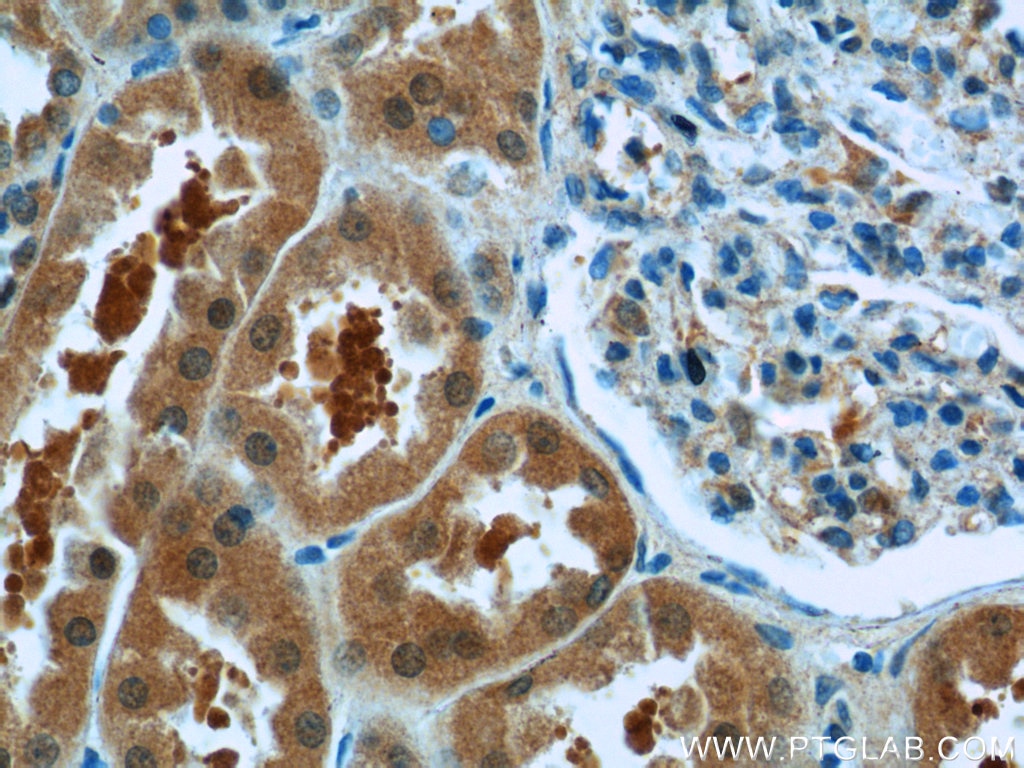
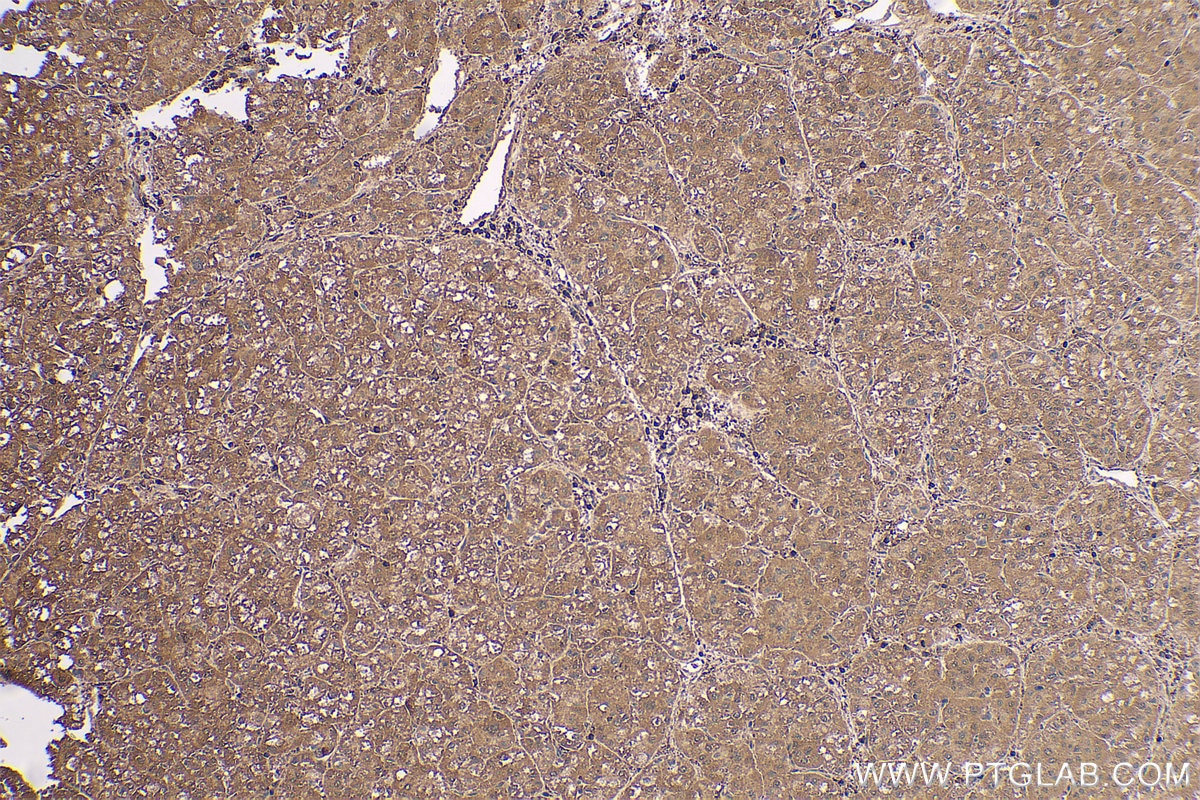
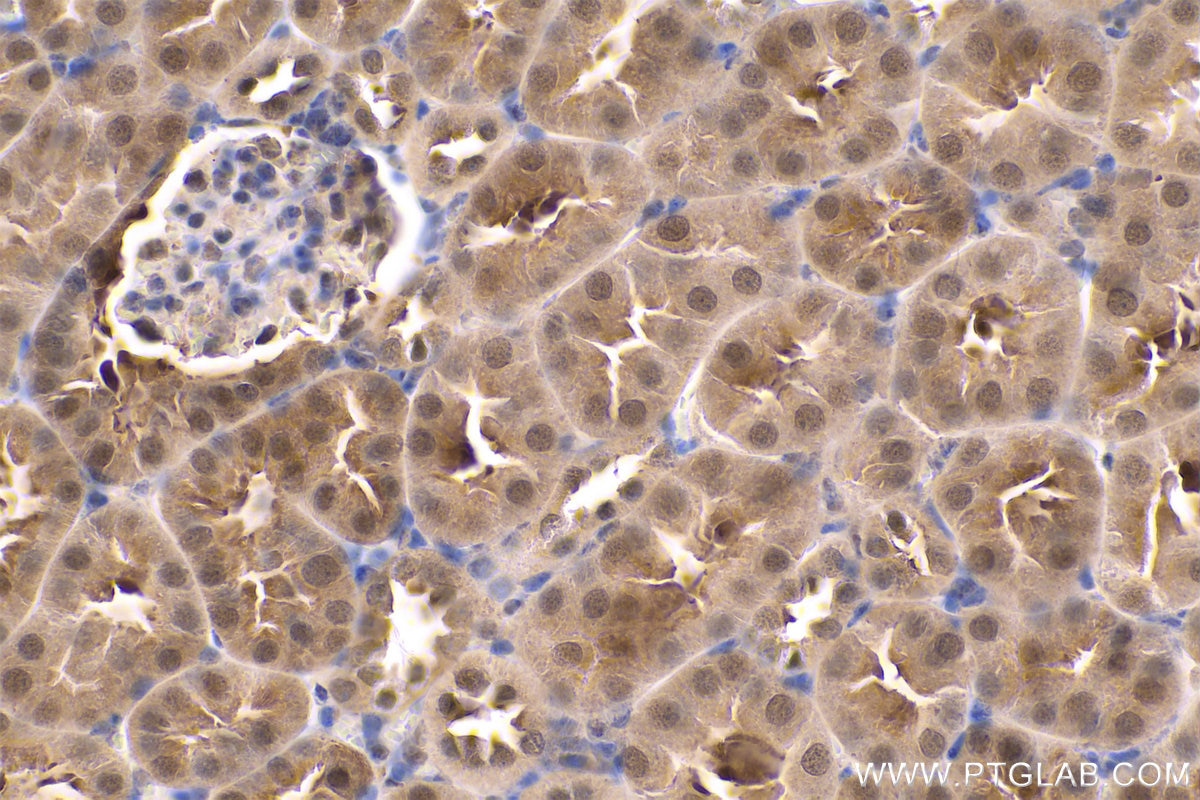
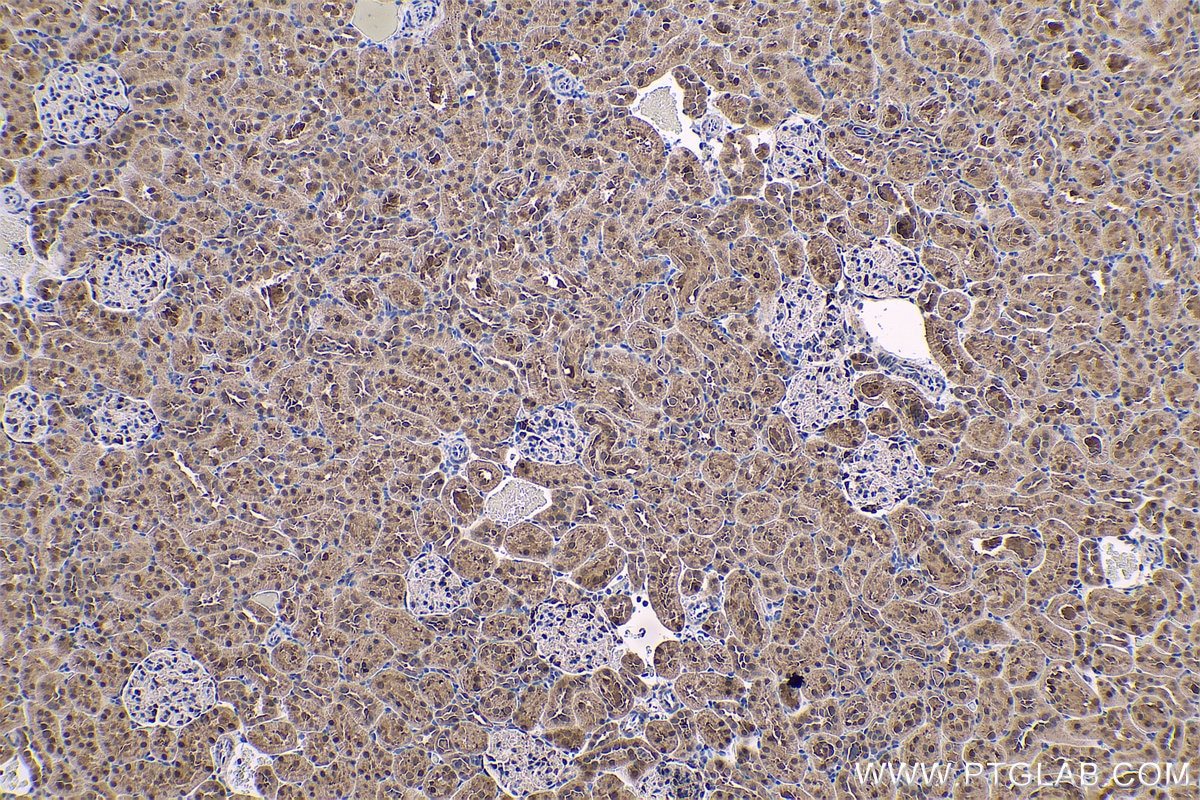
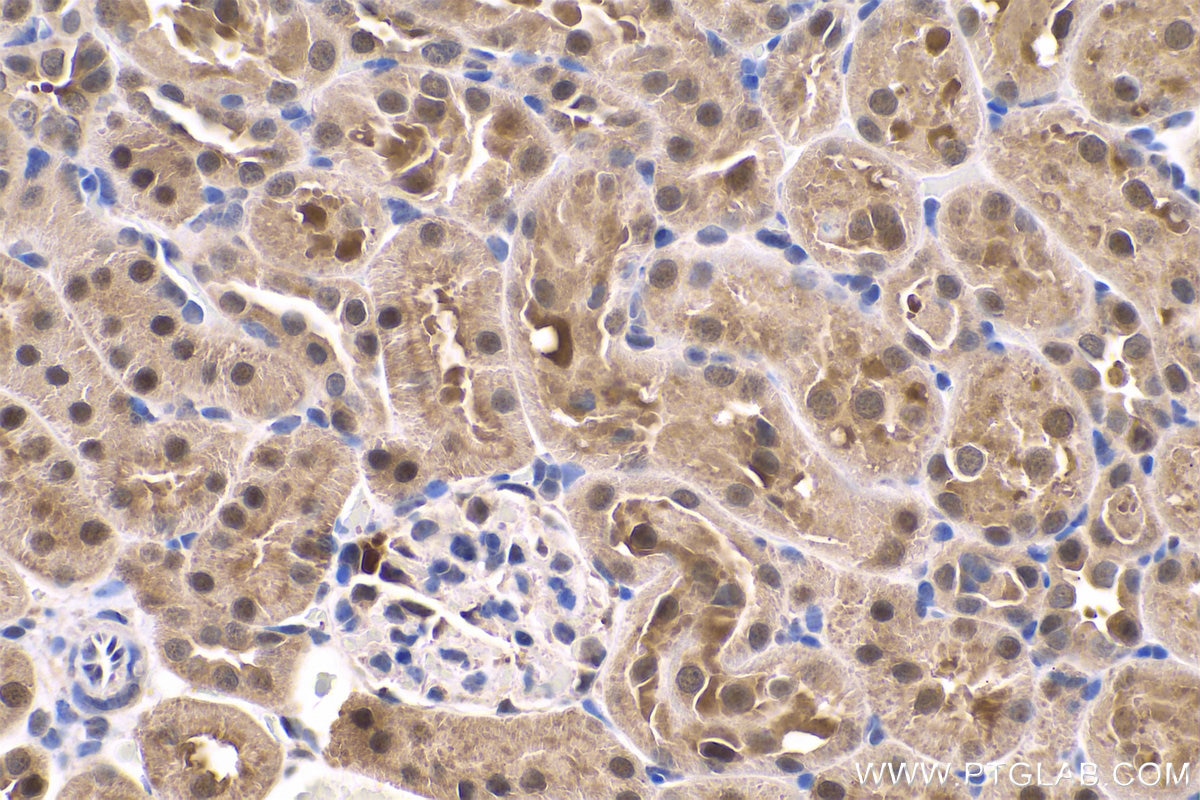
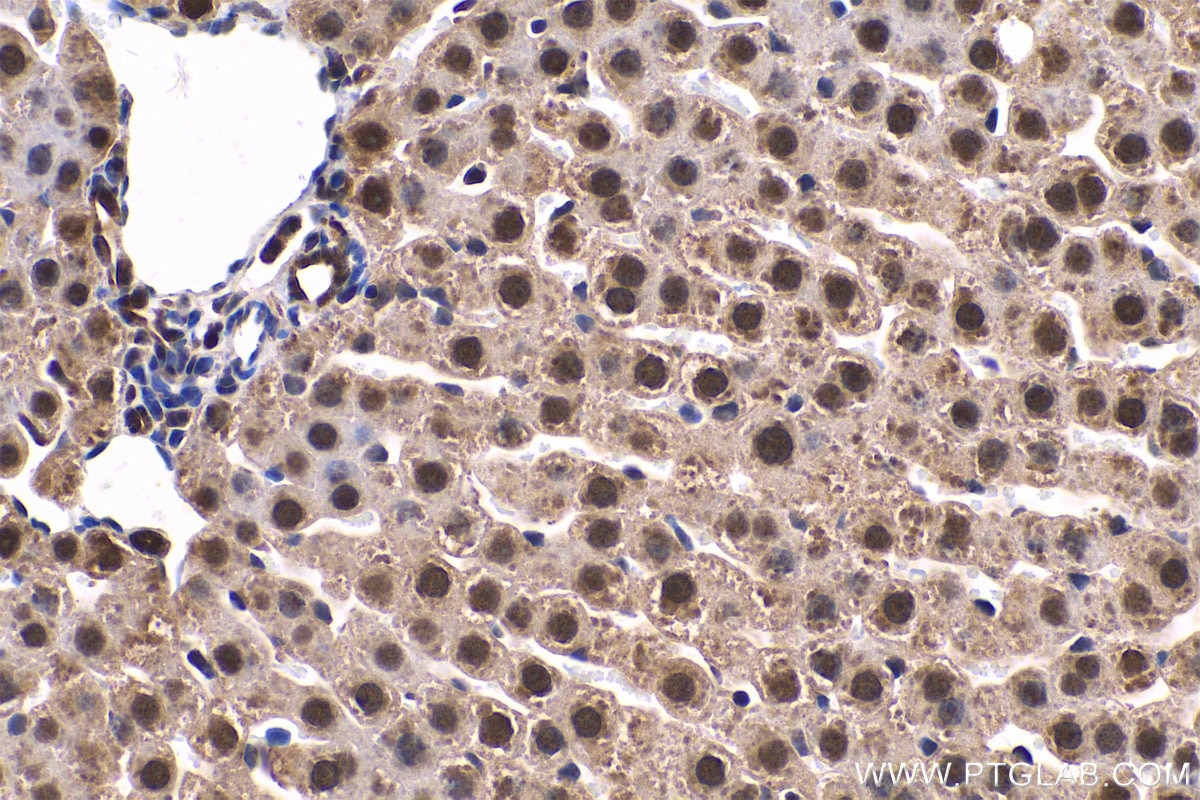
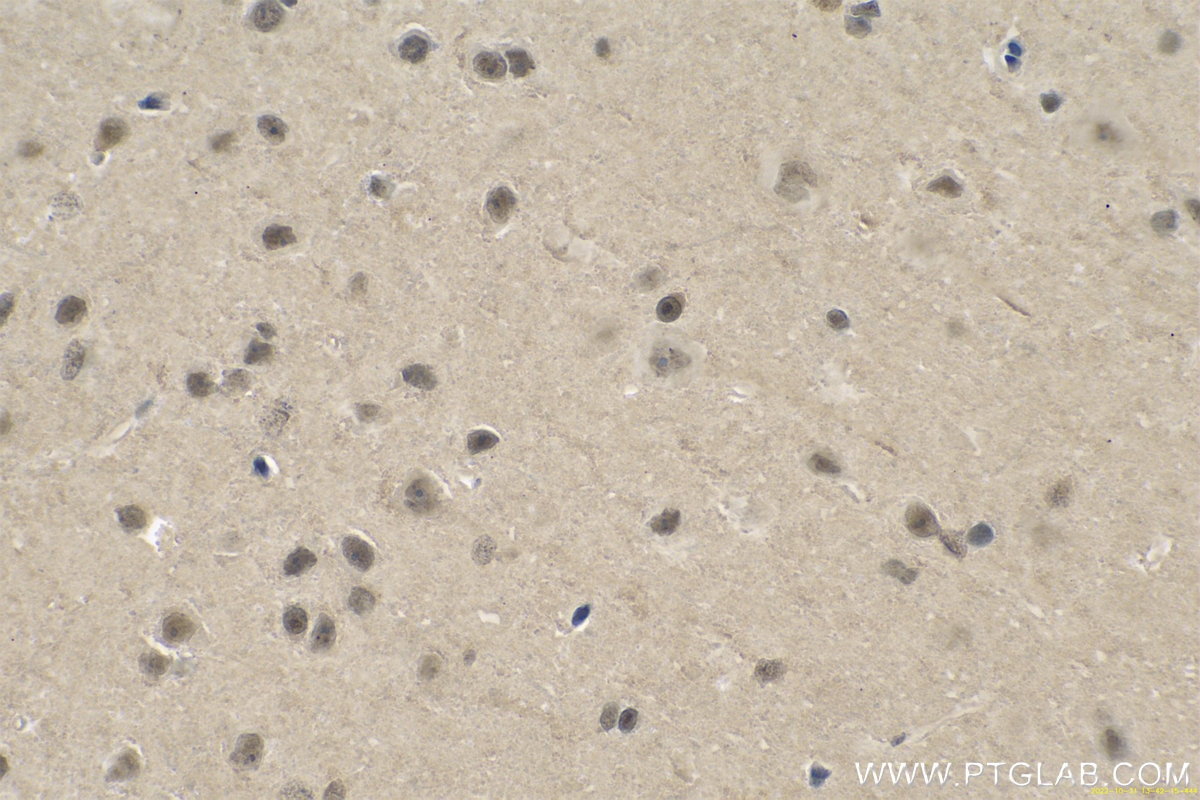
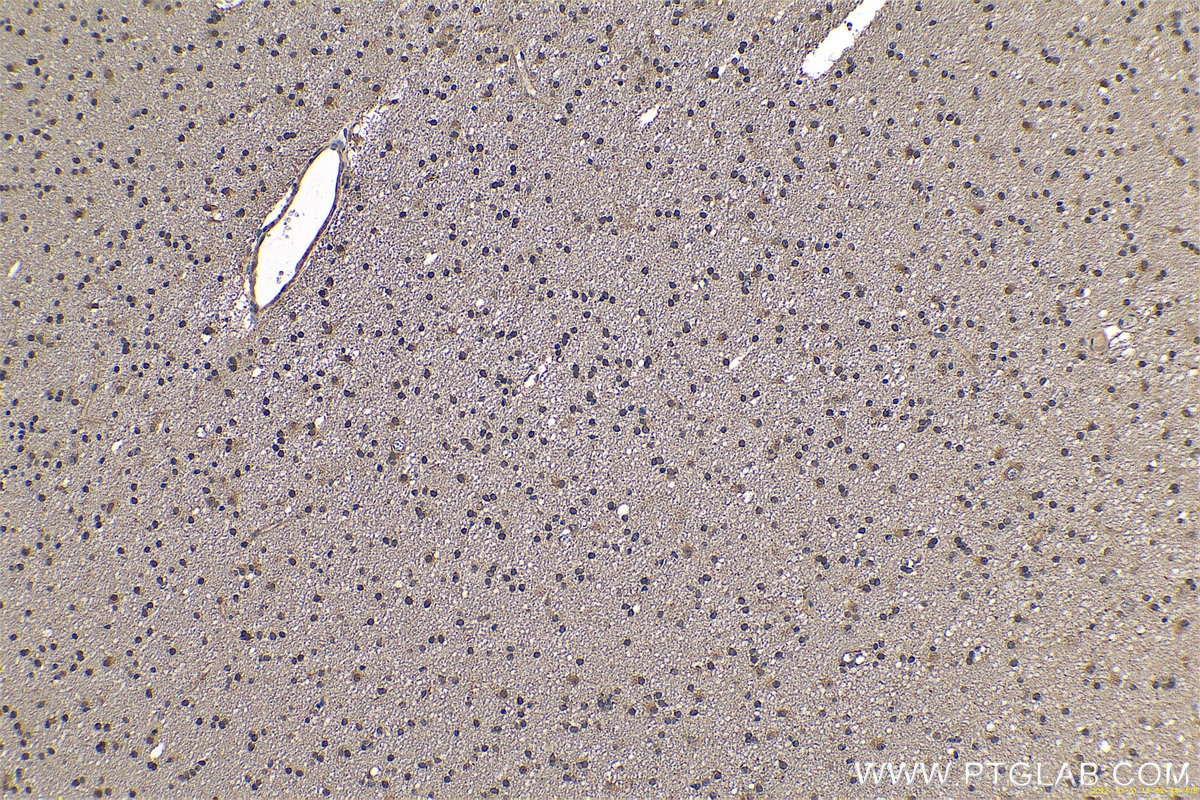
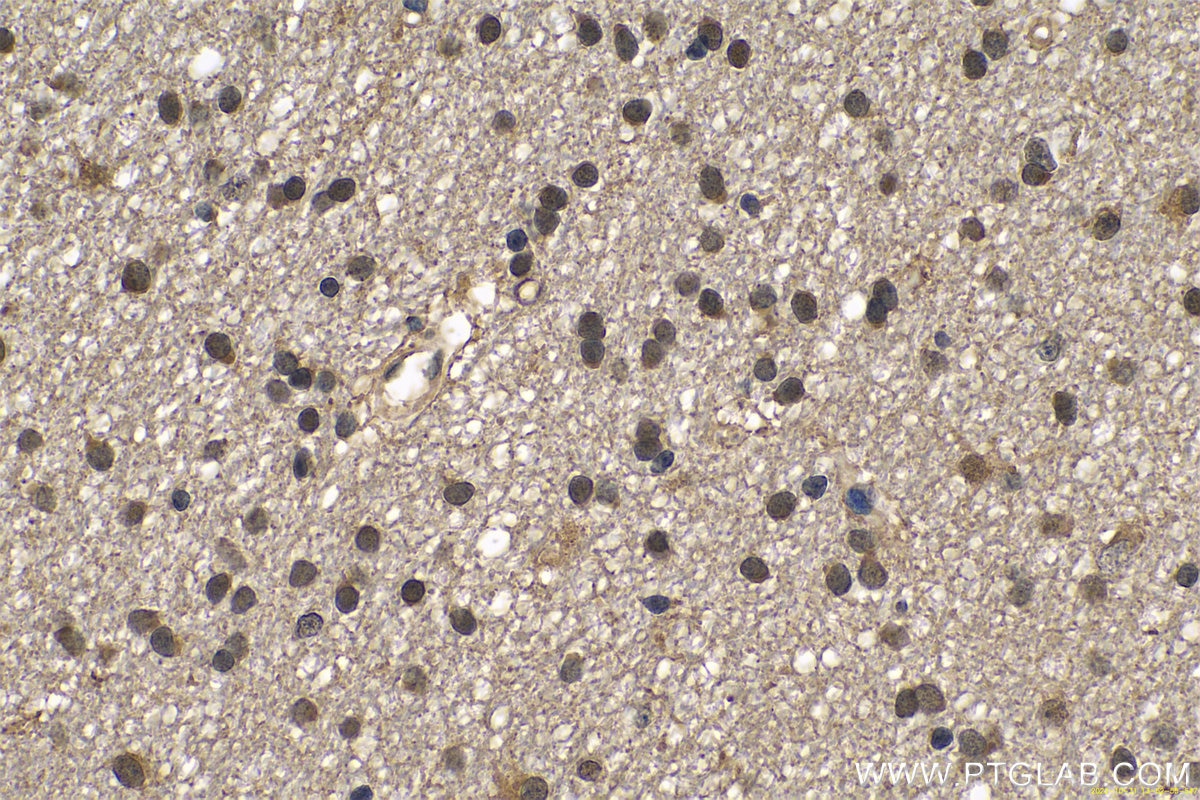
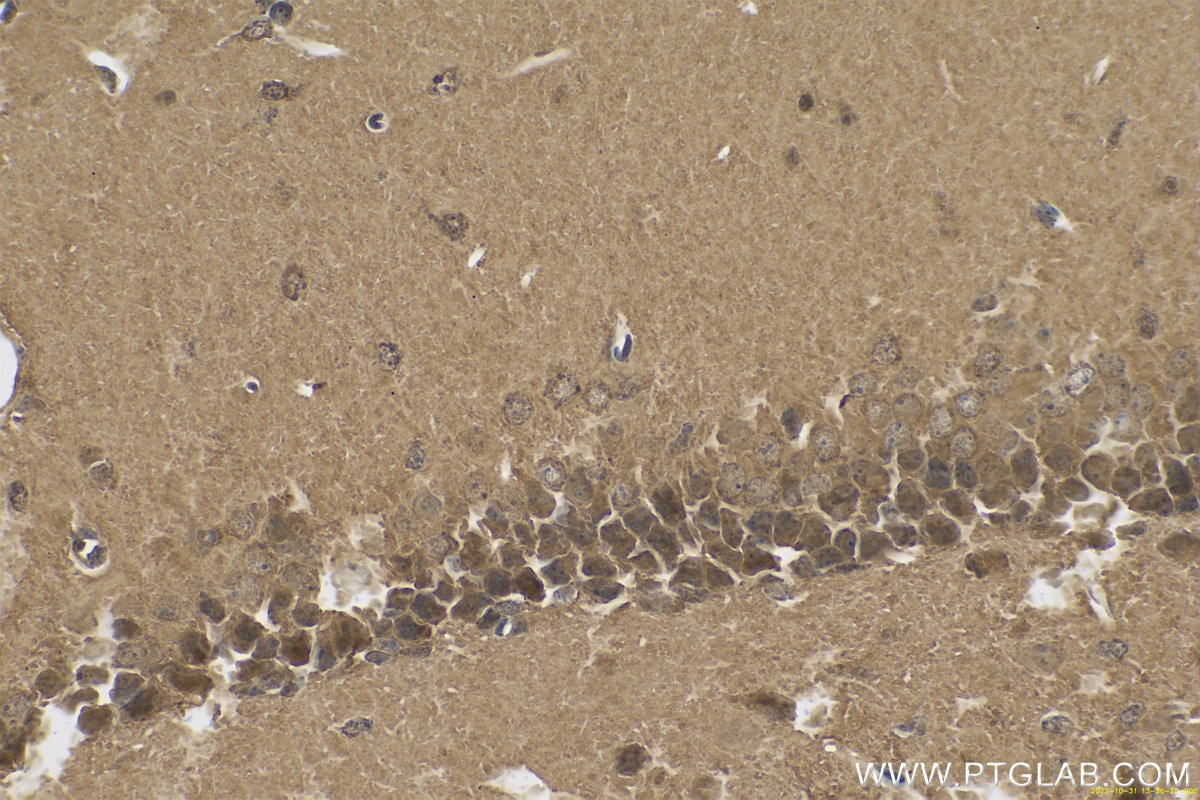
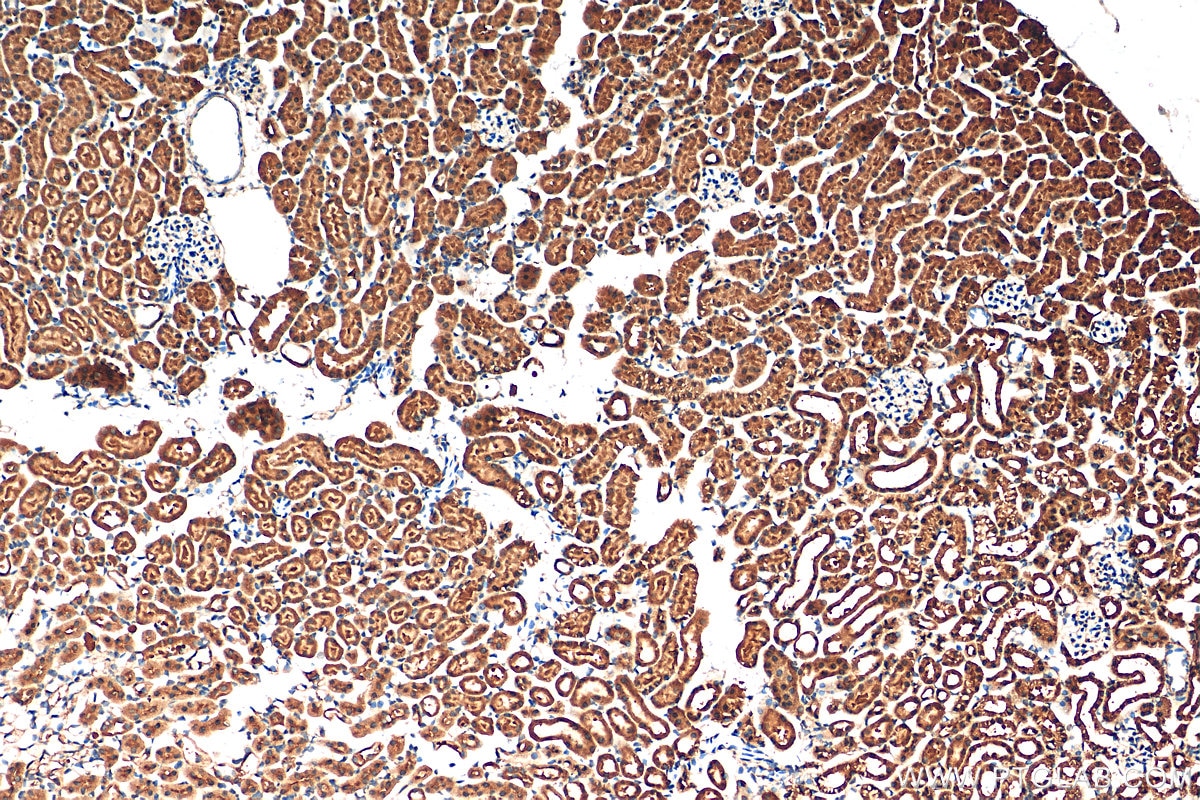
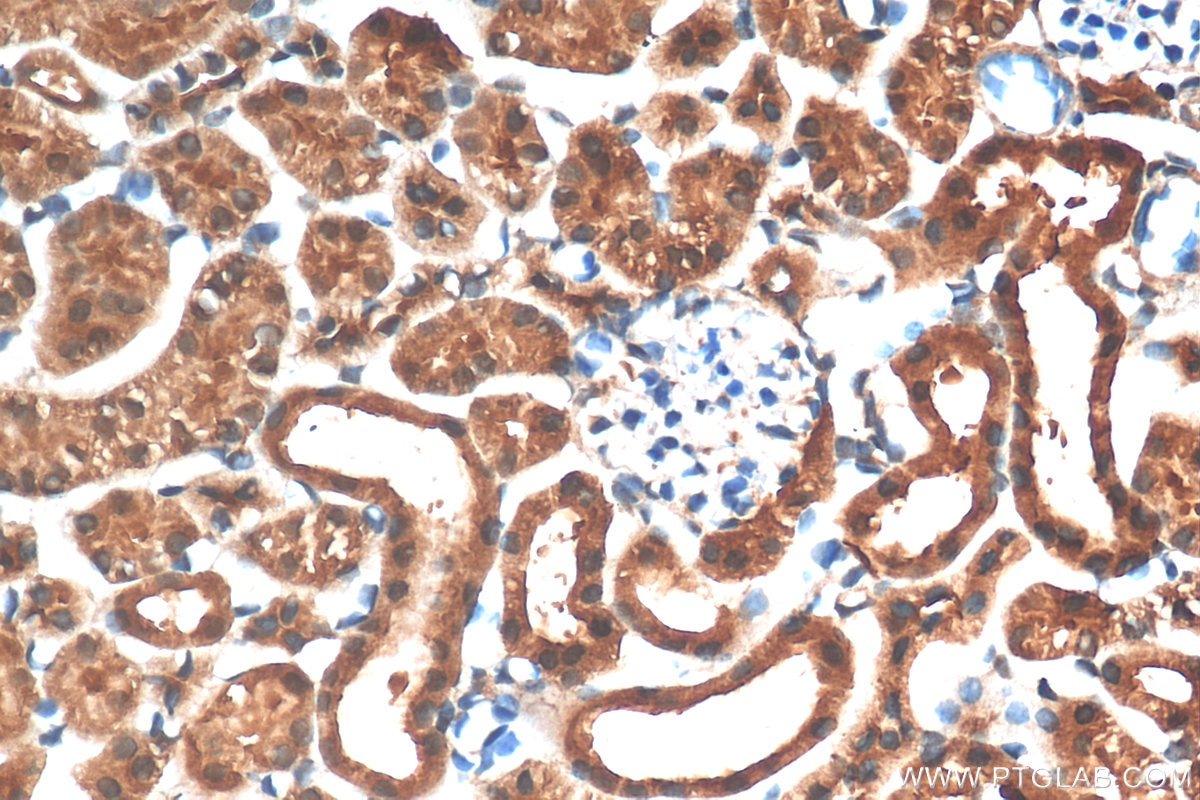
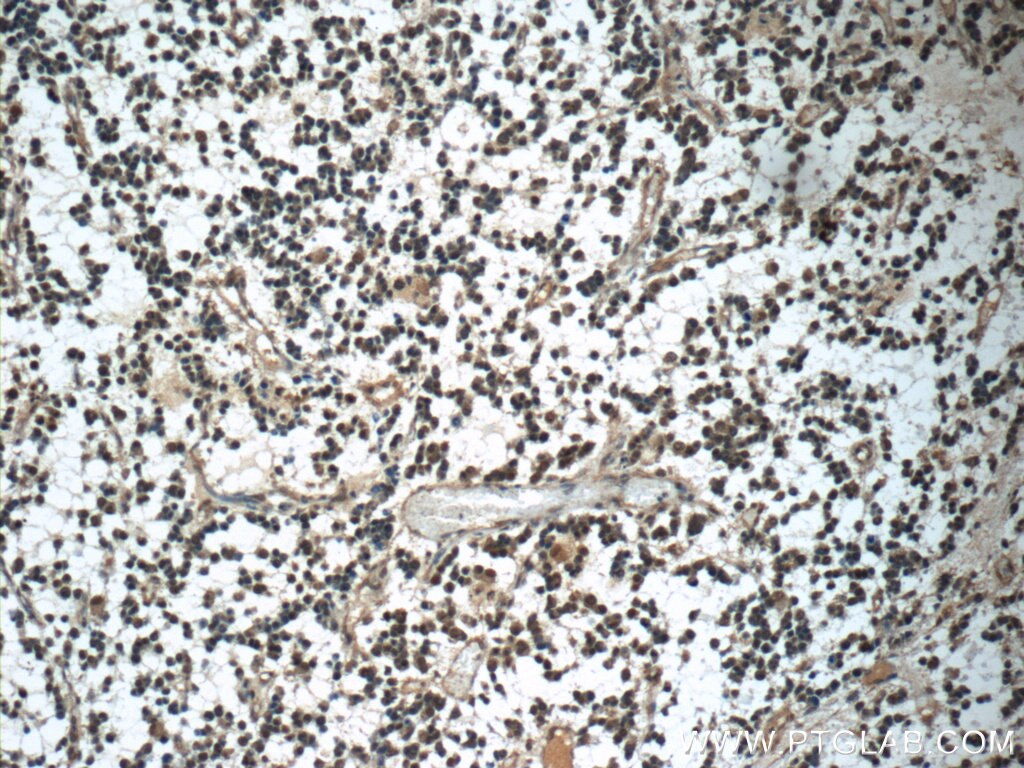
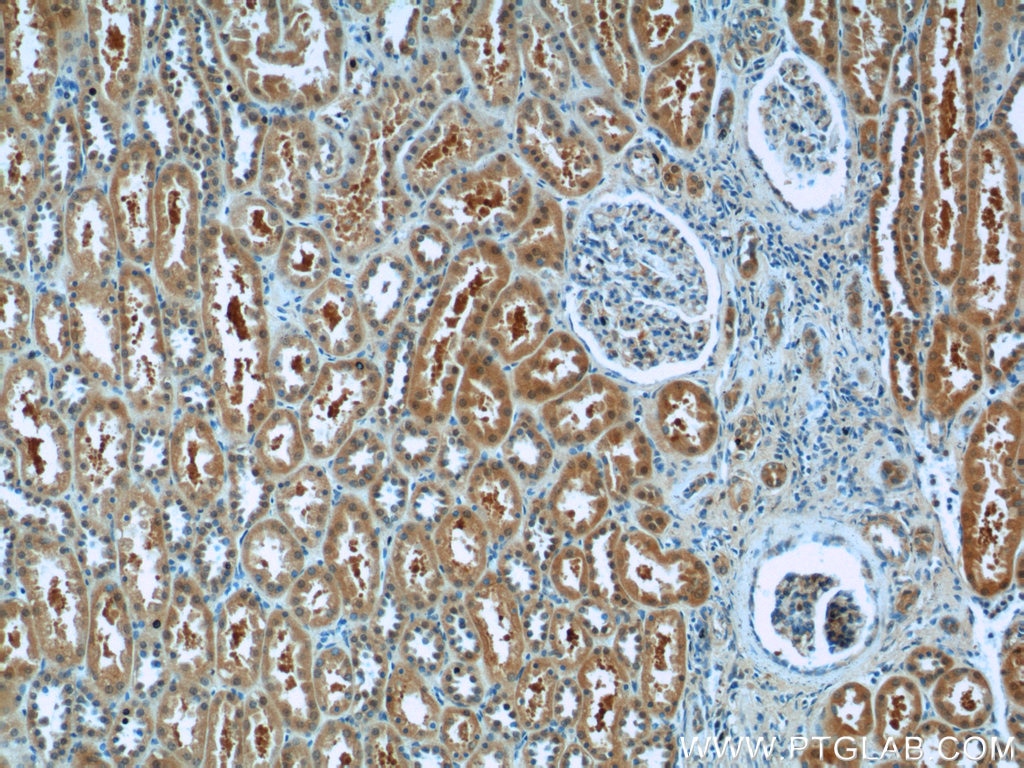
| Positive IHC detected in | human gliomas tissue, human liver cancer tissue, human kidney tissue, mouse kidney tissue, rat kidney tissue, rat liver tissue, mouse brain tissue, rat brain tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
| Positive IF/ICC detected in | SH-SY5Y cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:500-1:2000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Immunohistochemistry (IHC) | IHC : 1:1000-1:4000 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:50-1:500 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
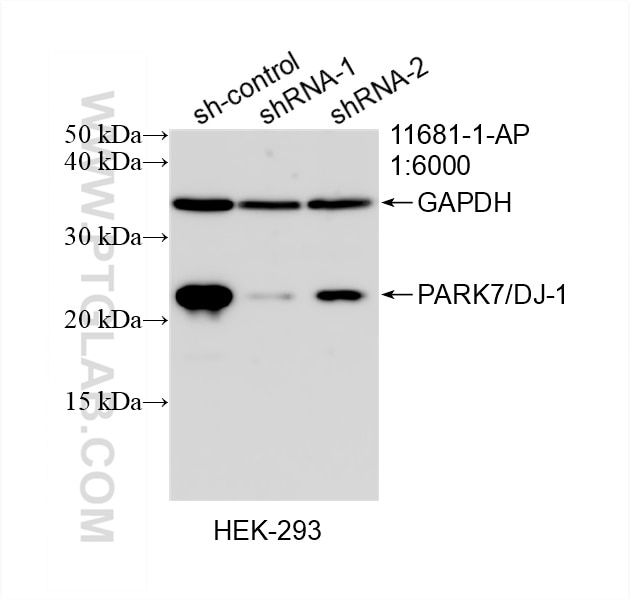
| KD/KO | See 3 publications below |
| WB | See 19 publications below |
| IHC | See 2 publications below |
| IF | See 3 publications below |
| IP | See 2 publications below |
Product Information
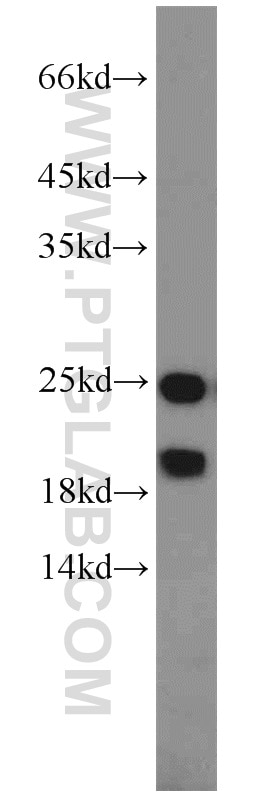
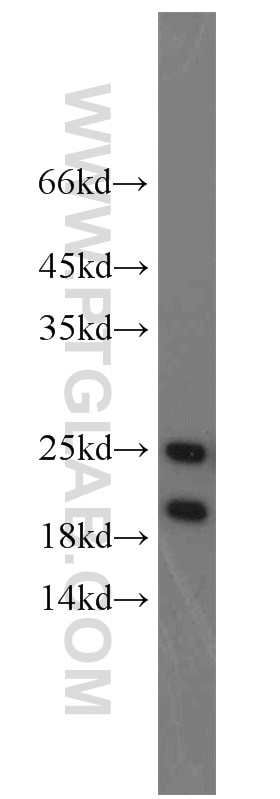
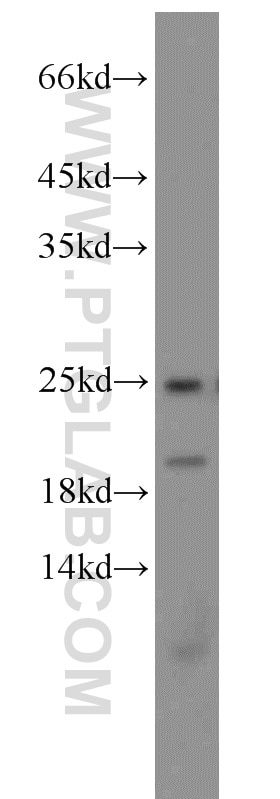
11681-1-AP targets PARK7/DJ-1 in WB, IHC, IF/ICC, IP, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag2287 Product name: Recombinant human PARK7,DJ-1 protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 1-189 aa of BC008188 Sequence: MASKRALVILAKGAEEMETVIPVDVMRRAGIKVTVAGLAGKDPVQCSRDVVICPDASLEDAKKEGPYDVVVLPGGNLGAQNLSESAAVKEILKEQENRKGLIAAICAGPTALLAHEIGFGSKVTTHPLAKDKMMNGGHYTYSENRVEKDGLILTSRGPGTSFEFALAIVEALNGKEVAAQVKAPLVLKD Predict reactive species |
| Full Name | Parkinson disease (autosomal recessive, early onset) 7 |
| Calculated Molecular Weight | 189 aa, 20 kDa |
| Observed Molecular Weight | 20 kDa, 25 kDa |
| GenBank Accession Number | BC008188 |
| Gene Symbol | PARK7 |
| Gene ID (NCBI) | 11315 |
| RRID | AB_2160099 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q99497 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
PARK7, also named as DJ1, belongs to the peptidase C56 family. It protects cells against oxidative stress and cell death. PARK7 plays a role in regulating expression or stability of the mitochondrial uncoupling proteins SLC25A14 and SLC25A27 in dopaminergic neurons of the substantia nigra pars compacta and attenuates the oxidative stress induced by calcium entry into the neurons via L-type channels during pacemaking. It eliminates hydrogen peroxide and protects cells against hydrogen peroxide-induced cell death. PARK7 has cell-growth promoting activity and transforming activity. It may function as a redox-sensitive chaperone. It's precursor undergoes a cleavage of a C-terminal peptide and subsequent activation of protease activity in response to oxidative stress. The amino acid replace at 166 (L → P) reduces PARK7 protein stability and leads to increased degradation. The predicted MW of this protein is 20 kDa, An additional 25 kDa band can be observed due to modification (PMID: 31767755).
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for PARK7/DJ-1 antibody 11681-1-AP | Download protocol |
| IHC protocol for PARK7/DJ-1 antibody 11681-1-AP | Download protocol |
| IP protocol for PARK7/DJ-1 antibody 11681-1-AP | Download protocol |
| WB protocol for PARK7/DJ-1 antibody 11681-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Nat Chem Biol Chemical proteomics reveals new targets of cysteine sulfinic acid reductase. | ||
Nat Commun Disuse-associated loss of the protease LONP1 in muscle impairs mitochondrial function and causes reduced skeletal muscle mass and strength.
| ||
Mol Cell Proteomics Chemoproteomics Reveals Chemical Diversity and Dynamics of 4-Oxo-2-nonenal Modifications in Cells. | ||
Mol Cell Biol ROS-mediated DJ-1 monomerization modulates intracellular trafficking involving Karyopherin β2. | ||
J Cell Mol Med FKBP3 aggravates the malignant phenotype of diffuse large B-cell lymphoma by PARK7-mediated activation of Wnt/β-catenin signalling |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH X (Verified Customer) (07-11-2022) | A very good and clean antibody with the right molecular weight for WB.
 |
FH Daniel (Verified Customer) (04-16-2019) | Specific band in RCC4 cells
 |