Mapping the mitotic protein interactome of Drosophila embryos with nanobodies
Immunoprecipitation of Protein Complexes from Drosophila Embryos with GFP-Trap
Mitosis is a fundamental process in life and it’s no wonder that mitosis is one of the best studied phenomena in biology. However, despite decades of intensive research, scientists are still far from understanding the molecular details of mitosis and its regulation.
So far, most of the data has been obtained from cultured cells. Such cells can be easily manipulated but have the drawback that they behave quite differently from the cells in an organism. On the other hand, in living organisms only a fraction of cells undergo mitosis at each point in time making it difficult to study this dynamic process. And still worse, research has been severely hampered by the lack of adequate tools to analyze (presumably short-lived and unstable) protein complexes thought to be involved in regulating mitosis.
A recently published methods paper by Lipinszki et al. (Cell Cycle Control: Mechanisms and Protocols, Methods in Molecular Biology vol. 1170: 571-588 (2014) describes an elegant genetic-proteomic approach to unravel the intricate molecular interactions that control mitosis. Here they exploit a special feature of the fruit fly Drosophila: Fertilized Drosophila eggs undergo a series of rapid and synchronous mitotic divisions of nuclei in a syncytium before cellularization of the blastoderm occurs. Divisions occur approx. every 10 min, whereby M and S phase alternate without intervening gap phases. Therefore, embryos collected before 2 h of age give a starting material highly enriched in mitotic protein complexes.
To be able to purify such complexes from Drosophila embryo extracts, Lipinszki et al. construct transgenic flies expressing GFP tagged baits (chimeric versions of endogenous proteins known to play a role in mitosis). Since the over-expressed chimeric proteins might interfere with oogenesis and/or early embryogenesis, they place their genes under the control of the inducible GAL4-UASp transcriptional regulator.
The decisive part of the experiment is pulling down GFP tagged protein complexes and analyzing the bound fraction by mass spectroscopy (MS).
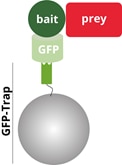
Yield, purity and speed are highly critical factors when isolating unstable complexes. As the authors note, standard GFP antibodies are not a good choice for this purpose, since the large immunoglobulin complexes, as major contaminants, cause “peptide masking” during mass spectrometric analysis. This may lead to the loss of important information about potential interactors or post-synthetic modifications. Lipinszki et al. show that using GFP-Trap for immunoprecipitation overcomes this problem. Since the one-step isolation of GFP-tagged baits with GFP-Trap is fast and efficient, the identification of interacting proteins works very well.
Over the past years, thousands of transgenic flies expressing proteins fused to GFP were established in order to follow the localization and dynamics of the protein of interest by microscopy (fixed preparations or live imaging). It makes life easier that the same fly lines can also be used for one-step affinity purifications of the GFP fusion proteins and its interacting factors. Combined knowledge of localization and interactions is highly informative on a protein’s function. And quite obviously, this approach also bears potential to probe interactions involved in other processes that occur in Drosophila early embryos or other organisms.
Selected literature
- Lipinszki Z, Wang P, Grant R, Lindon C, Dzhindzhev NS, D’Avino PP, Przewloka MR, Glover DM and Archambault V Cell Cycle Control: Mechanisms and Protocols, Methods in Molecular Biology vol. 1170: 571-588 (2014)