Tested Applications
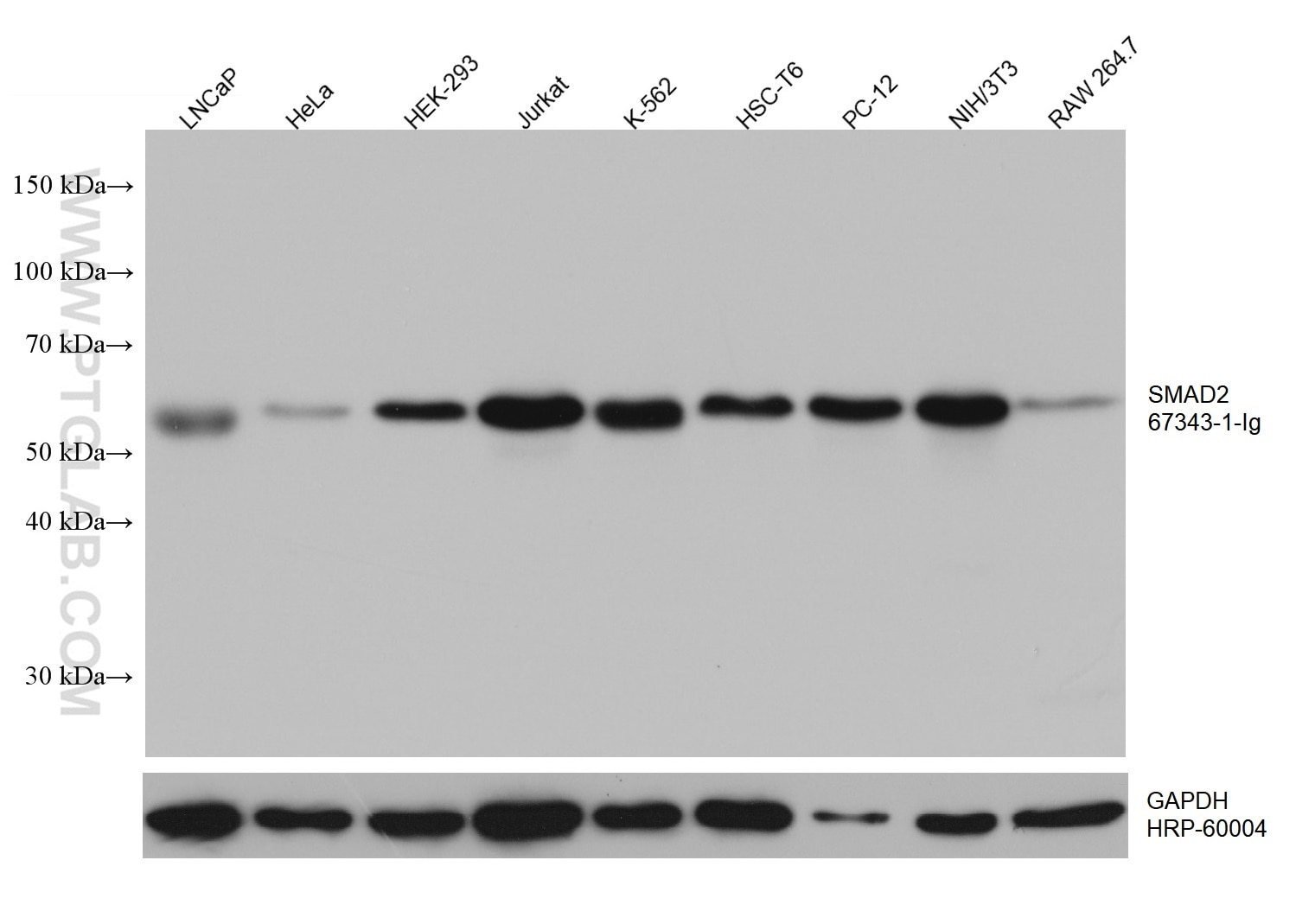
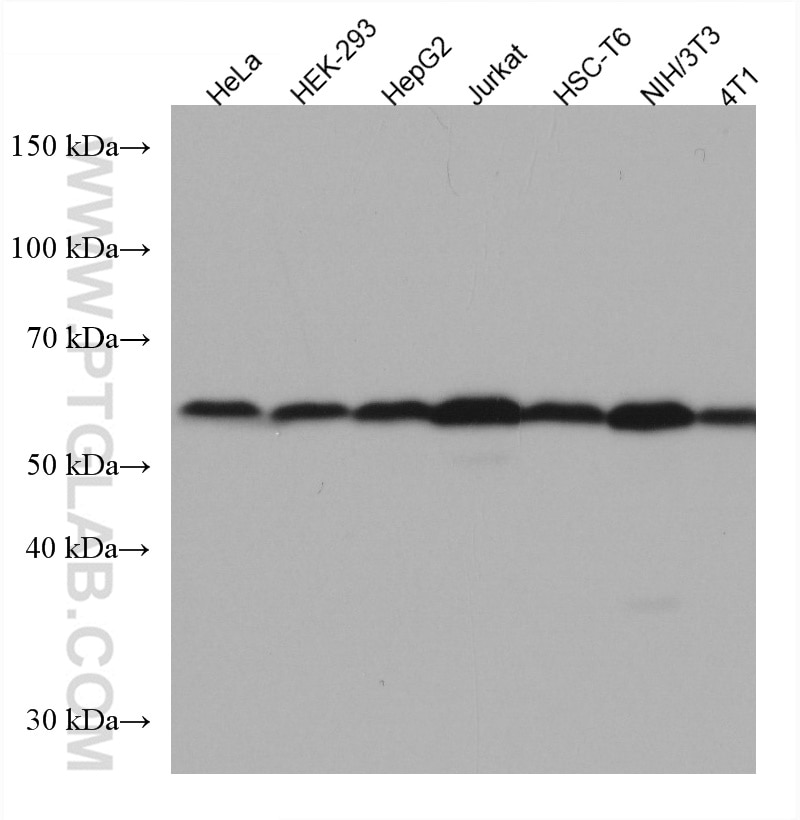
| Positive WB detected in | HeLa cells, LNCaP cells, HEK-293 cells, Jurkat cells, K-562 cells, HSC-T6 cells, PC-12 cells, NIH/3T3 cells, RAW 264.7 cells, HepG2 cells, 4T1 cells |
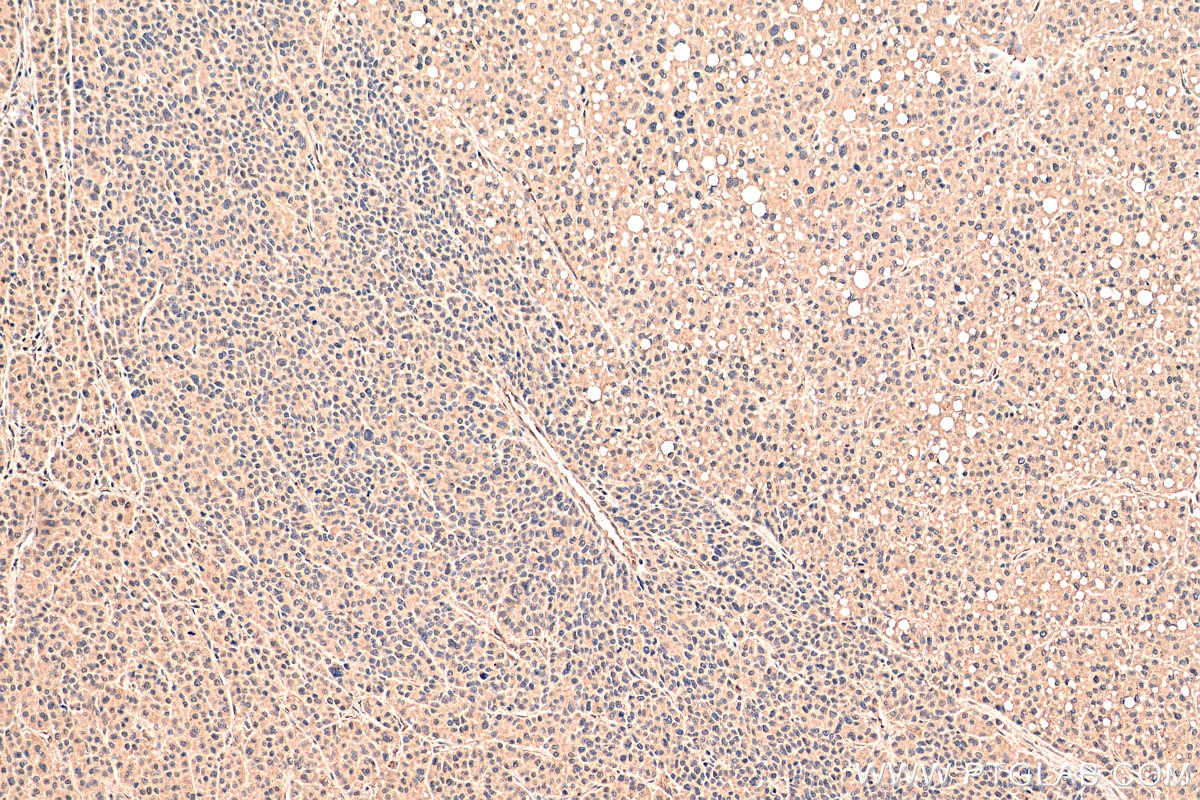
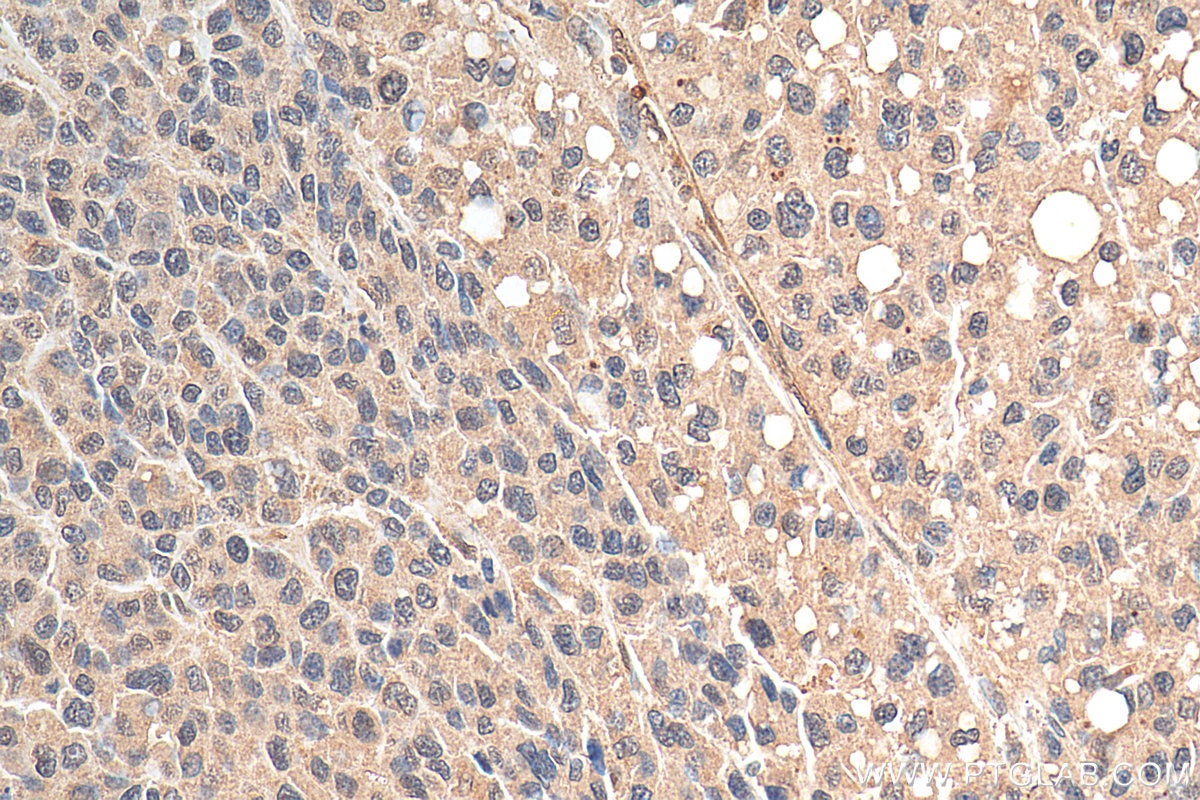
| Positive IHC detected in | human liver cancer tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
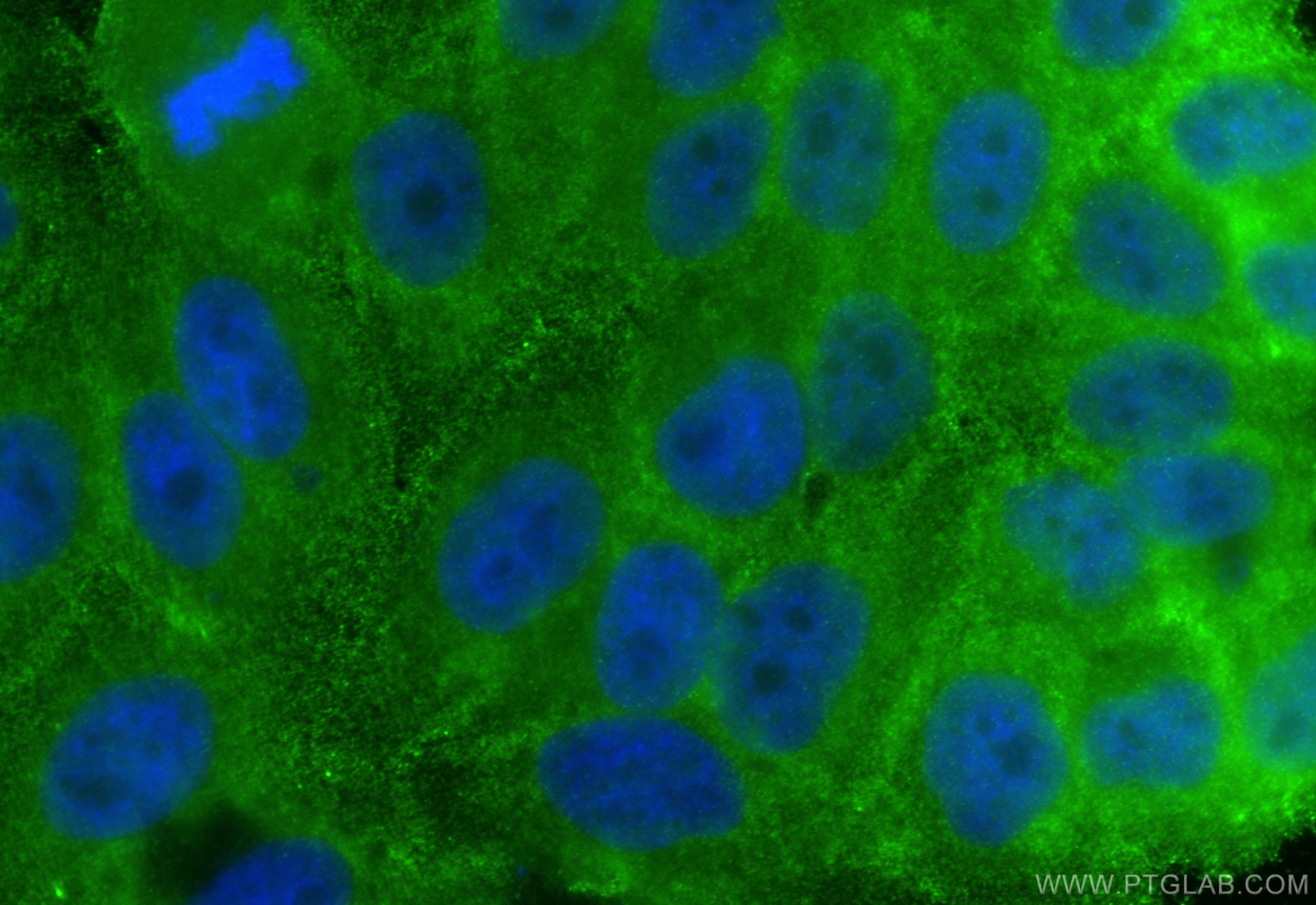
| Positive IF/ICC detected in | HepG2 cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:2000-1:10000 |
| Immunohistochemistry (IHC) | IHC : 1:1000-1:4000 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:400-1:1600 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| WB | See 17 publications below |
Product Information
67343-1-Ig targets SMAD2 in WB, IHC, IF/ICC, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Mouse / IgG1 |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag19542 Product name: Recombinant human SMAD2 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 1-355 aa of BC014840 Sequence: MSSILPFTPPVVKRLLGWKKSAGGSGGAGGGEQNGQEEKWCEKAVKSLVKKLKKTGRLDELEKAITTQNCNTKCVTIPSTCSEIWGLSTPNTIDQWDTTGLYSFSEQTRSLDGRLQVSHRKGLPHVIYCRLWRWPDLHSHHELKAIENCEYAFNLKKDEVCVNPYHYQRVETPVLPPVLVPRHTEILTELPPLDDYTHSIPENTNFPAGIEPQSNYIPETPPPGYISEDGETSDQQLNQSMDTGSPAELSPTTLSPVNHSLDLQPVTYSEPAFWCSIAYYELNQRVGETFHASQPSLTVDGFTDPSNSERFCLGLLSNVNRNATVEMTRRHIGRGVRLYYIGGEVFAECLSDSAI Predict reactive species |
| Full Name | SMAD family member 2 |
| Calculated Molecular Weight | 467 aa, 52 kDa |
| Observed Molecular Weight | 58 kDa |
| GenBank Accession Number | BC014840 |
| Gene Symbol | SMAD2 |
| Gene ID (NCBI) | 4087 |
| RRID | AB_2882601 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein G purification |
| UNIPROT ID | Q15796 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
SMAD2, also named as MADH2 and MADR2, belongs to the dwarfin/SMAD family, contains 1 MH1 (MAD homology 1) domain and 1 MH2 (MAD homology 2) domain. SMAD2 is a receptor-regulated SMAD(R-SMAD) that is an intracellular signal transducer and transcriptional modulator activated by TGF-beta and activin type 1 receptor kinases. This protein may act as a tumor suppressor in colorectal carcinoma. It is phosphorylated on one or several of Thr-220, Ser-245, Ser-250, and Ser-255. In response to TGF-beta, It is phosphorylated on Ser-465/467 by TGF-beta and activin type 1 receptor kinases, and then able to interact with SMURF2, recruiting other proteins, such as SNON, for degradation. In response to decorin, the naturally occurring inhibitor of TGF-beta signaling, it is phosphorylated on Ser-240 by CaMK2. It is phosphorylated by MAPK3 upon EGF stimulation; which increases transcriptional activity and stability, and is blocked by calmodulin. In response to TGF-beta, it is ubiquitinated by NEDD4L, which promotes its degradation. In response to TGF-beta signaling, it is acetylated on Lys-19 by coactivators, which increases transcriptional activity. The molecular weight of unphosphorylated forms of Smad2 is 52 kDa and phosphorylated forms of Smad2 is 58 kDa. (PMID: 9006934)
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for SMAD2 antibody 67343-1-Ig | Download protocol |
| IHC protocol for SMAD2 antibody 67343-1-Ig | Download protocol |
| WB protocol for SMAD2 antibody 67343-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
J Cell Mol Med Mesenchymal stem cells ameliorate silica-induced pulmonary fibrosis by inhibition of inflammation and epithelial-mesenchymal transition. | ||
Pediatr Pulmonol Effect of electro-acupuncture at ST 36 on maternal food restriction-induced lung phenotype in rat offspring. | ||
RSC Adv Quercetin ameliorates podocyte injury via inhibition of oxidative stress and the TGF-β1/Smad pathway in DN rats | ||
Evid Based Complement Alternat Med The Traditional Chinese Medicine Formula FTZ Protects against Cardiac Fibrosis by Suppressing the TGFβ1-Smad2/3 Pathway. | ||
J Agric Food Chem Biosynthesis of the Natural Antioxidant Mycosporine-Glycine and Its Anti-Photodamage Properties | ||
Tissue Cell ADAM17 promotes colorectal cancer migration and invasion by regulating the TGF-β/Smad signaling pathway |