Tested Applications
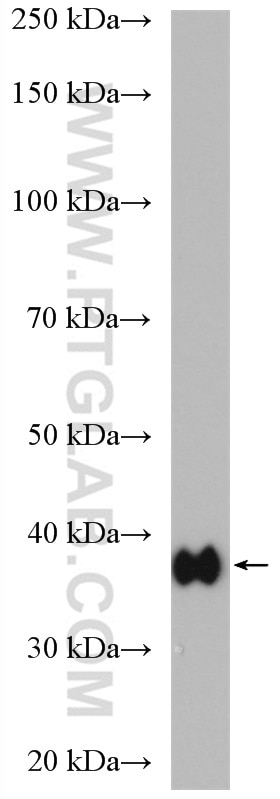
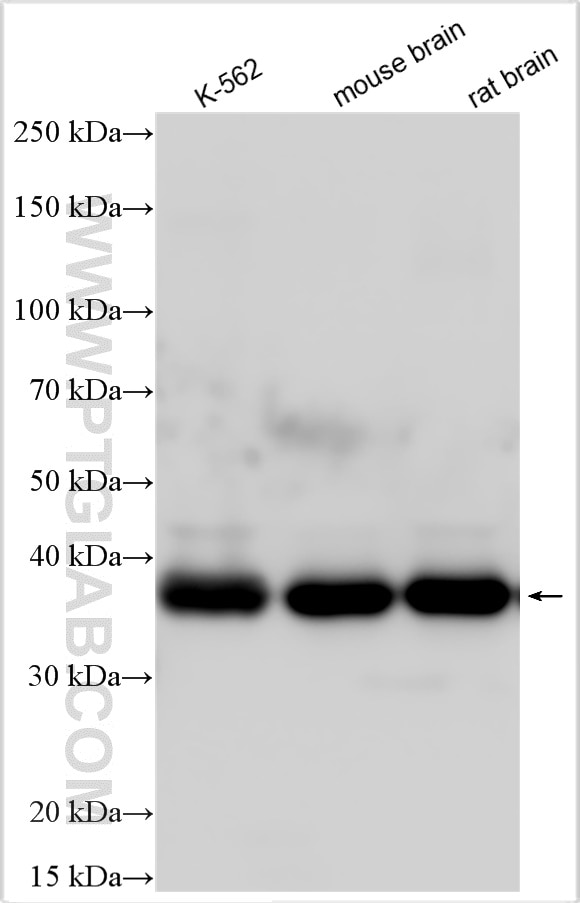
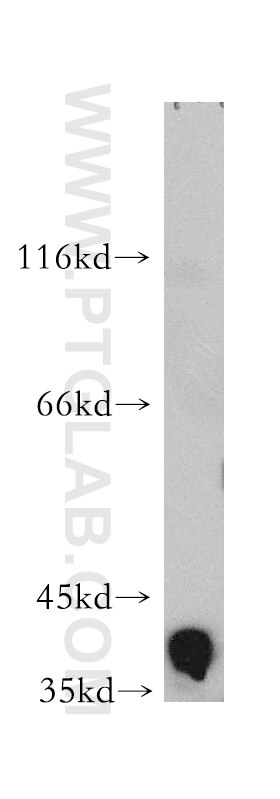
| Positive WB detected in | K-562 cells, human heart tissue, mouse cerebellum tissue |
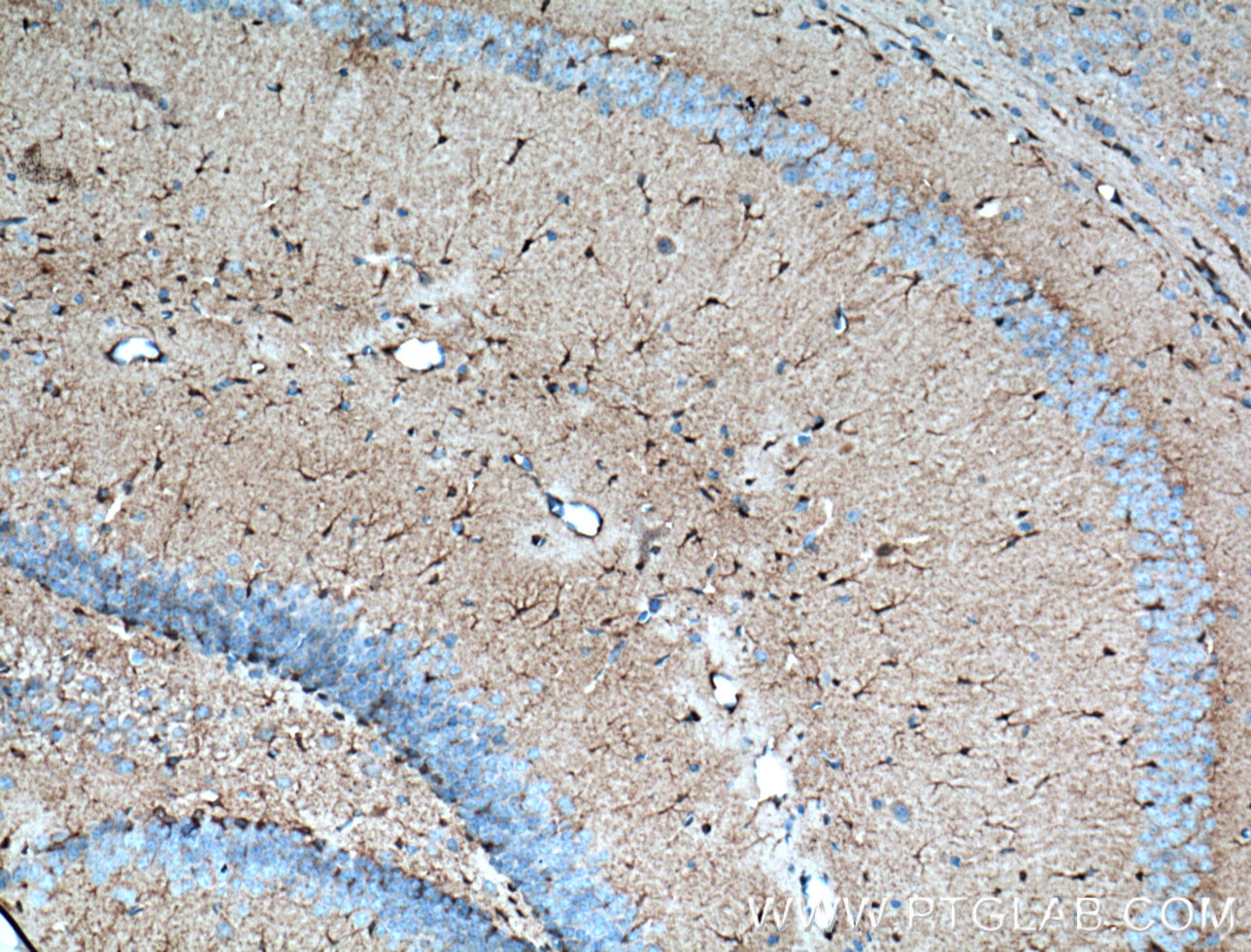
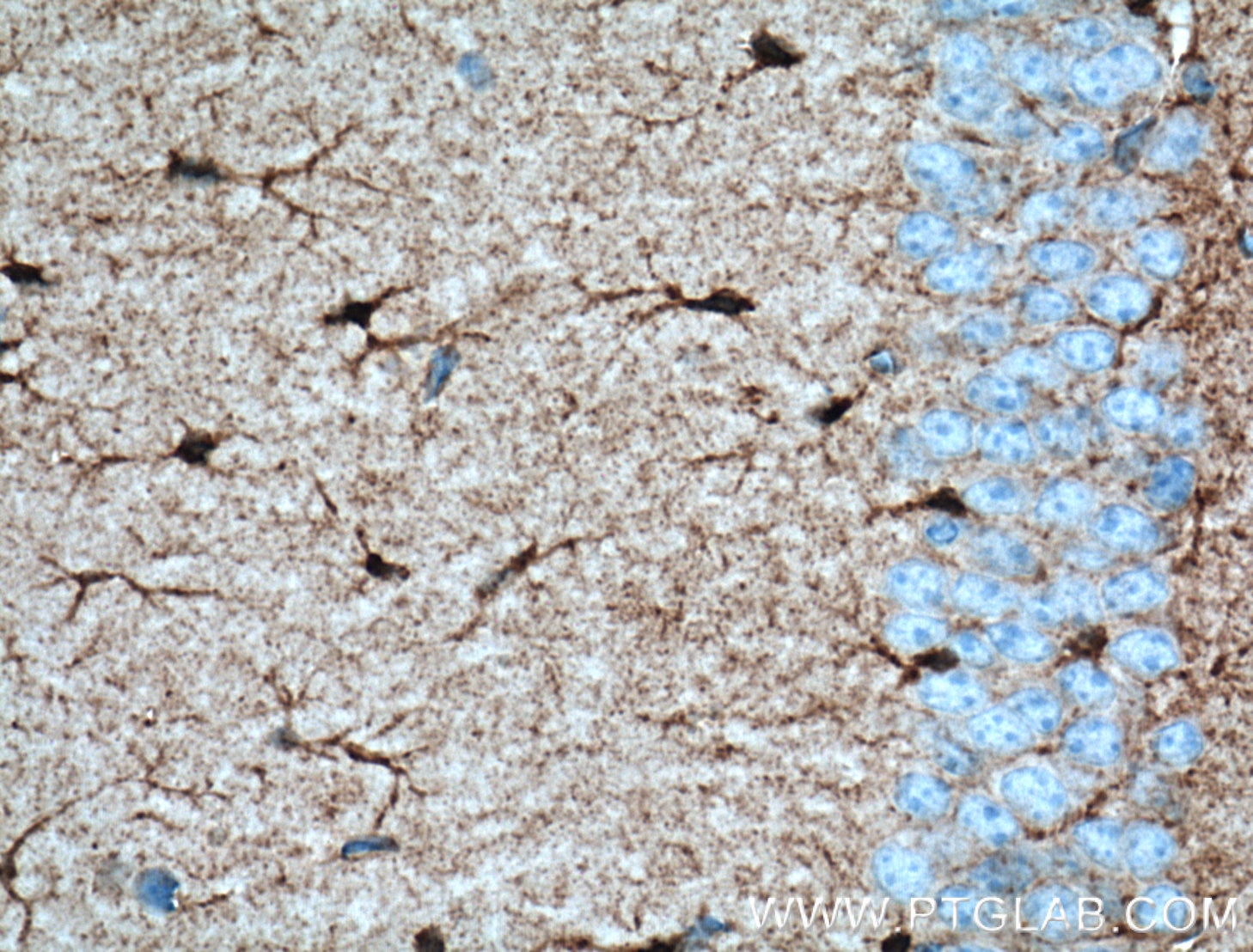
| Positive IHC detected in | mouse brain tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
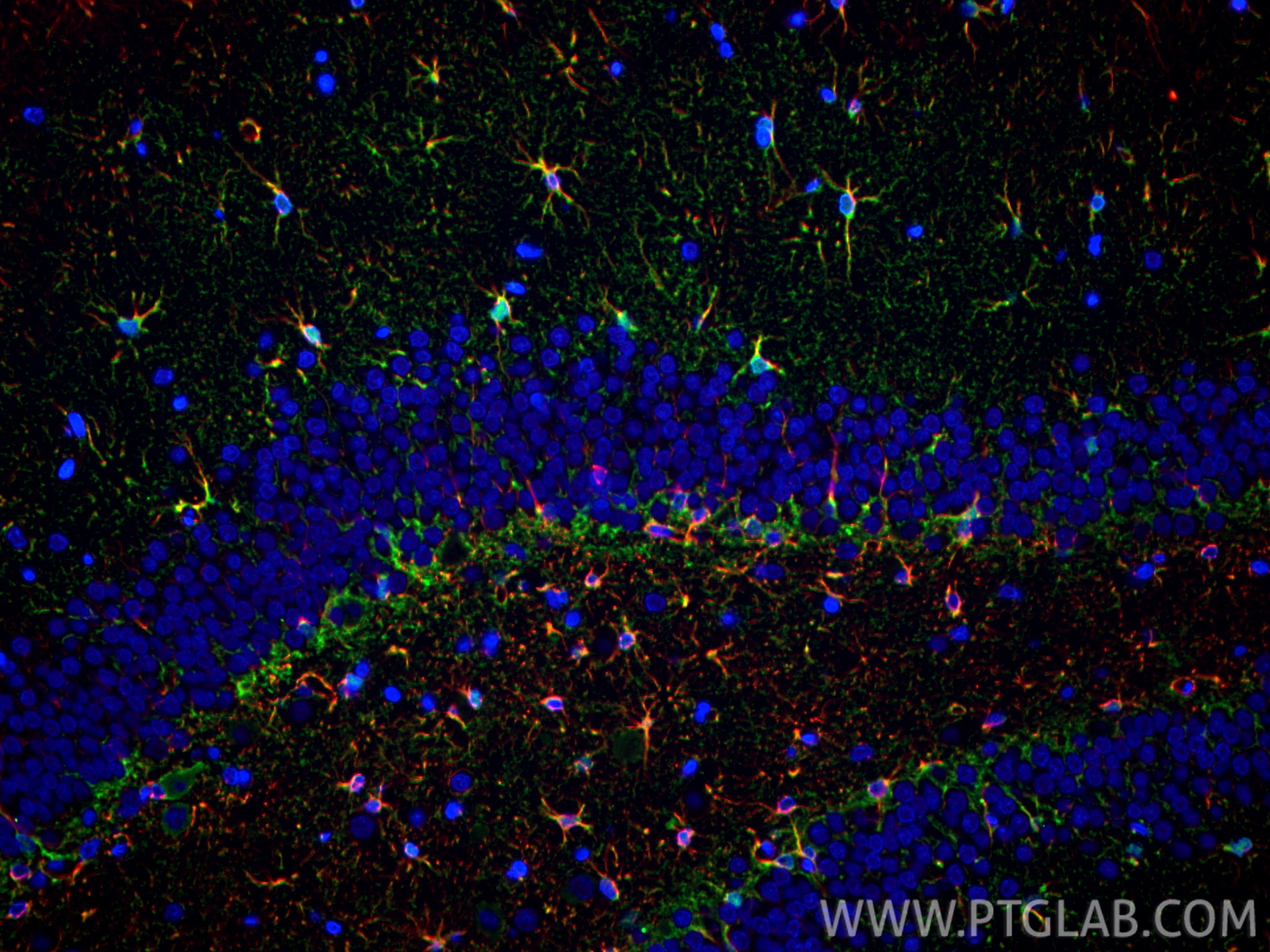
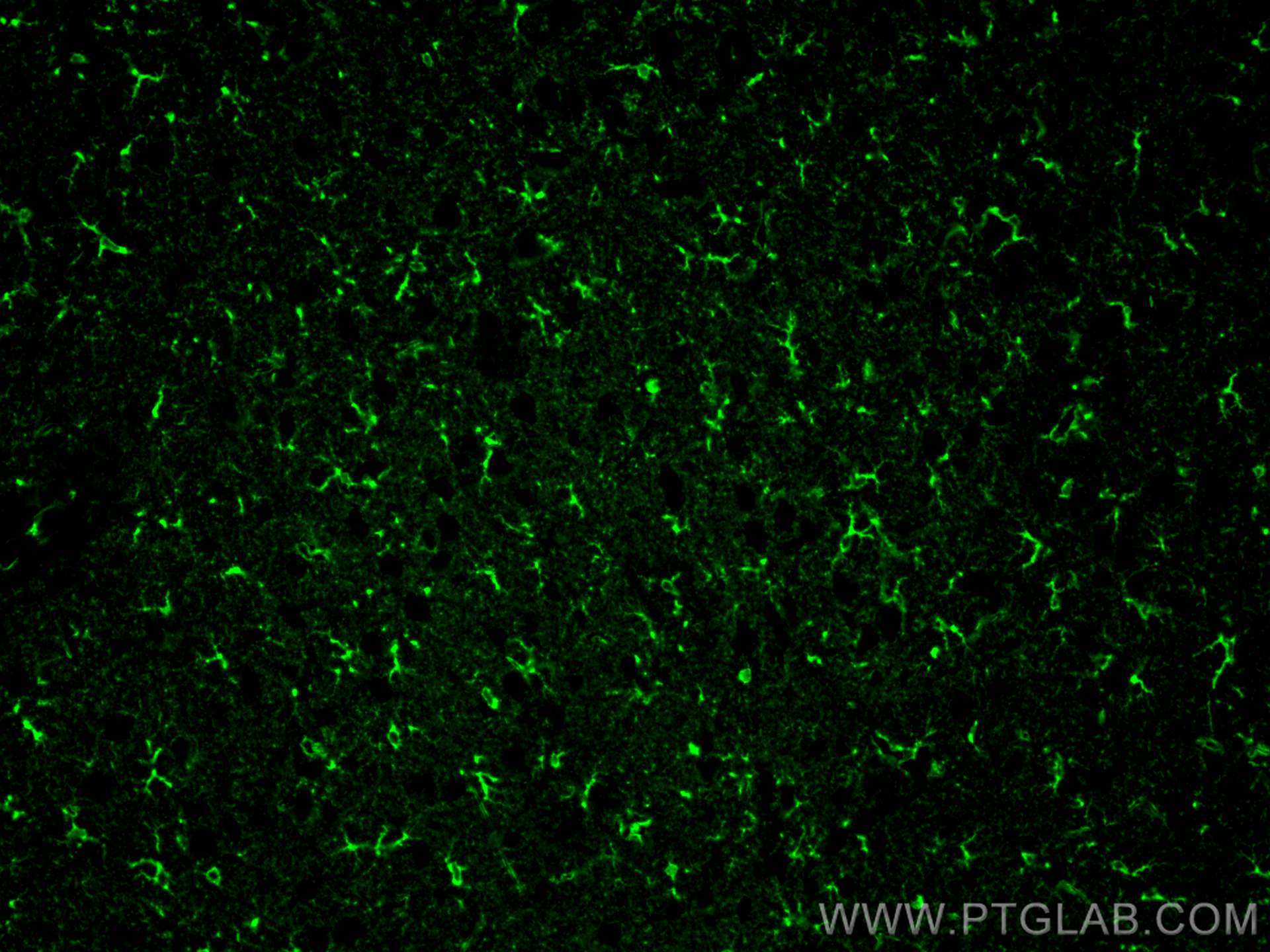
| Positive IF-P detected in | mouse brain tissue, rat brain tissue |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:5000-1:50000 |
| Immunohistochemistry (IHC) | IHC : 1:50-1:500 |
| Immunofluorescence (IF)-P | IF-P : 1:200-1:800 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 1 publications below |
| WB | See 22 publications below |
| IHC | See 3 publications below |
| IF | See 3 publications below |
| IP | See 1 publications below |
| ELISA | See 1 publications below |
Product Information
14884-1-AP targets Aldolase C in WB, IHC, IF-P, IP, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse, chicken |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag6659 Product name: Recombinant human ALDOC protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 1-364 aa of BC003613 Sequence: MPHSYPALSAEQKKELSDIALRIVAPGKGILAADESVGSMAKRLSQIGVENTEENRRLYRQVLFSADDRVKKCIGGVIFFHETLYQKDDNGVPFVRTIQDKGIVVGIKVDKGVVPLAGTDGETTTQGLDGLSERCAQYKKDGADFAKWRCVLKISERTPSALAILENANVLARYASICQQNGIVPIVEPEILPDGDHDLKRCQYVTEKVLAAVYKALSDHHVYLEGTLLKPNMVTPGHACPIKYTPEEIAMATVTALRRTVPPAVPGVTFLSGGQSEEEASFNLNAINRCPLPRPWALTFSYGRALQASALNAWRGQRDNAGAATEEFIKRAEVNGLAAQGKYEGSGEDGGAAAQSLYIANHAY Predict reactive species |
| Full Name | aldolase C, fructose-bisphosphate |
| Calculated Molecular Weight | 39 kDa |
| Observed Molecular Weight | 39 kDa |
| GenBank Accession Number | BC003613 |
| Gene Symbol | Aldolase C |
| Gene ID (NCBI) | 230 |
| RRID | AB_2226691 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | P09972 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
Fructose-bisphosphate aldolase C (ALDOC) reversibly cleaves FBP and F1-P to glyceraldehyde 3-phosphate and dihydroxyacetone phosphate and is strongly expressed in mammalian brain together with ALDOA and is alsopresent in the heart and spleen of some species(PMID:9363598). It is involved in glycolysis as an important enzyme. Phospholipase D2 and inositol 1,4,5-triphosphate interact with ALDOC in signal transduction. Meanwhile, the protein expression of ALDOC has been reported to be regulated in brain tumor, hepatomas, and lung cancer(PMID:21548097).14884-1-AP can recognize Aldolase A and Aldolase C
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for Aldolase C antibody 14884-1-AP | Download protocol |
| IHC protocol for Aldolase C antibody 14884-1-AP | Download protocol |
| WB protocol for Aldolase C antibody 14884-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Nat Commun O-GlcNAc modification of leucyl-tRNA synthetase 1 integrates leucine and glucose availability to regulate mTORC1 and the metabolic fate of leucine. | ||
Nat Commun A network of RNA-binding proteins controls translation efficiency to activate anaerobic metabolism. | ||
Brain C9orf72 expansion within astrocytes reduces metabolic flexibility in amyotrophic lateral sclerosis. | ||
Mol Ther Single-Cell Transcriptome Analysis Reveals Intratumoral Heterogeneity in ccRCC, which Results in Different Clinical Outcomes. | ||
Hum Mol Genet Lack of CUL4B leads to increased abundance of GFAP-positive cells that is mediated by PTGDS in mouse brain. | ||
Stem Cell Reports The Long Noncoding RNA Lncenc1 Maintains Naive States of Mouse ESCs by Promoting the Glycolysis Pathway. |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH Kazu (Verified Customer) (11-22-2022) | This antibody worked on frozen mouse brain tissue sections. A good astrocyte marker.
|