iCLIP for the thorough analysis of mRNA:protein interactions
Individual-nucleotide resolution UV crosslinking and immunoprecipitation (iCLIP) of GFP-tagged RNA-binding proteins with GFP-Trap
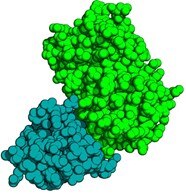
Crystal structure of the anti-GFP VHH-Green Fluorescent Protein complex.
The GFP Nanobody is displayed blue and the GFP in green color.
UV crosslinking techniques are the method of choice for a comprehensive analysis of in-vivo-mRNA targets of an RNA-binding protein (RBP). In the recent publication of Olgeiser et al. (2019), the authors applied individual-nucleotide resolution UV crosslinking and immunoprecipitation (iCLIP) to study fungal mRNA transport. For this approach, they have used strains expressing GFP-tagged versions of the two RBPs Grp1 and Rrm4; an optimized protocol was developed to uncover that Grp1 and Rrm4 conjointly bind thousands of shared target messenger ribonucleoproteins (mRNPs) in the fungus U. maydis. The protein:RNA complexes were immunoprecipitated in a multiple detergent containing buffer using ChromoTek’s GFP-Trap Magnetic Agarose. This is a transcriptome‐wide view to an endosomal mRNA transport machinery.
The detailed protocol is given in the recent publications “The key protein of endosomal mRNP transport Rrm4 binds translational landmark sites of cargo mRNAs” by Olgeiser et al. in EMBO reports (2019) 20: e46588 and “Adaption of iCLIP to plants determines the binding landscape of the clock-regulated RNA-binding protein AtGRP7” by Meyer et al in Genome Biology (2017) 18:204.