Tested Applications
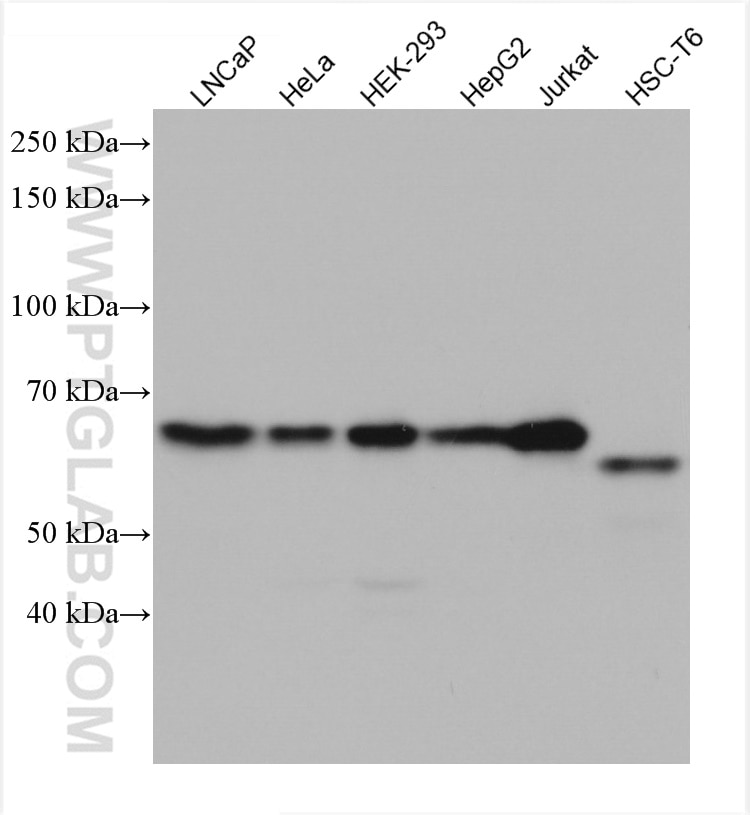
| Positive WB detected in | LNCaP cells, HepG2 cells, HeLa cells, HEK-293 cells, Jurkat cells, HSC-T6 cells |
Recommended dilution
| Application | Dilution |
|---|---|
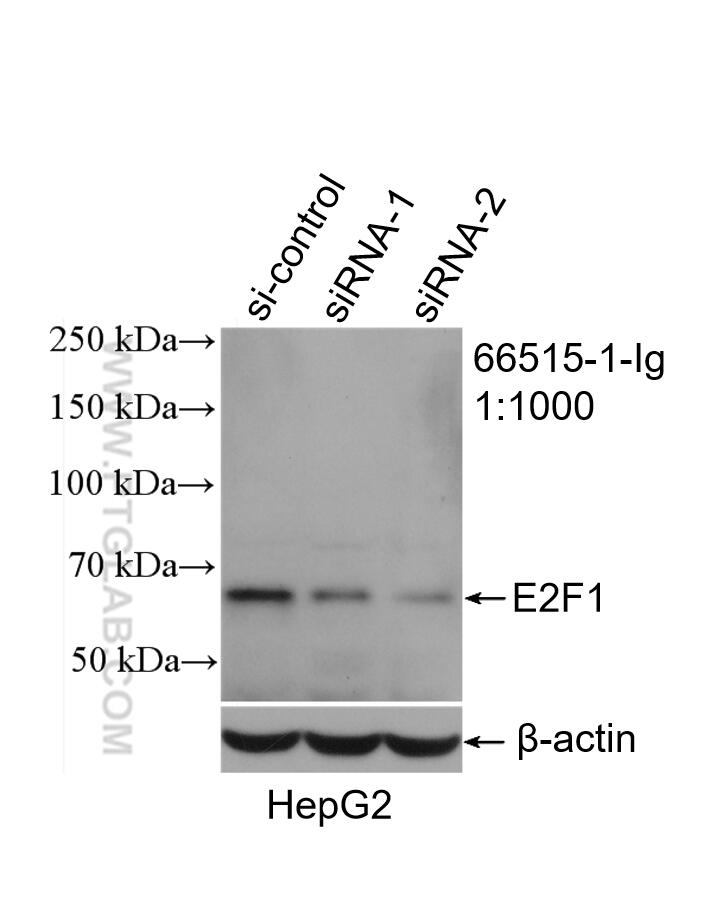
| Western Blot (WB) | WB : 1:1000-1:6000 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 16 publications below |
| WB | See 74 publications below |
| IHC | See 10 publications below |
| IF | See 8 publications below |
| IP | See 4 publications below |
| CoIP | See 5 publications below |
| ChIP | See 17 publications below |
Product Information
66515-1-Ig targets E2F1 in WB, IHC, IF, IP, CoIP, ChIP, ELISA applications and shows reactivity with human, rat samples.
| Tested Reactivity | human, rat |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Mouse / IgG2b |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag17363 Product name: Recombinant human E2F1 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 88-438 aa of BC050369 Sequence: VKRRLDLETDHQYLAESSGPARGRGRHPGKGVKSPGEKSRYETSLNLTTKRFLELLSHSADGVVDLNWAAEVLKVQKRRIYDITNVLEGIQLIAKKSKNHIQWLGSHTTVGVGGRLEGLTQDLRQLQESEQQLDHLMNICTTQLRLLSEDTDSQRLAYVTCQDLRSIADPAEQMVMVIKAPPETQLQAVDSSENFQISLKSKQGPIDVFLCPEETVGGISPGKTPSQEVTSEEENRATDSATIVSPPPSSPPSSLTTDPSQSLLSLEQEPLLSRMGSLRAPVDEDRLSPLVAADSLLEHVREDFSGLLPEEFISLSPPHEALDYHFGLEEGEGIRDLFDCDFGDLTPLDF Predict reactive species |
| Full Name | E2F transcription factor 1 |
| Calculated Molecular Weight | 437 aa, 47 kDa |
| Observed Molecular Weight | 55-60 kDa |
| GenBank Accession Number | BC050369 |
| Gene Symbol | E2F1 |
| Gene ID (NCBI) | 1869 |
| RRID | AB_2881878 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein A purification |
| UNIPROT ID | Q01094 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
Transcription factor E2F1 (E2F1), also known as RBBP3, is a transcription activator that binds DNA cooperatively with dp proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The DRTF1/E2F complex functions in the control of cell-cycle progression from G1 to S phase. E2F-1 binds preferentially RB1 protein, in a cell-cycle dependent manner. It can mediate both cell proliferation and p53-dependent apoptosis. The calculated molecular weight of E2F1 is 47 kDa, but the sumoylated E2F1 is bout 55-60 kDa.
Protocols
| Product Specific Protocols | |
|---|---|
| WB protocol for E2F1 antibody 66515-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cancer Res LncRNA AGPG confers endocrine resistance in breast cancer by promoting E2F1 activity | ||
J Adv Res Transducin-like enhancer of split 3 protects against lipopolysaccharide-induced inflammation through DDX5-ATF1-PPP2R5A signaling | ||
Cell Death Dis ZNF652 exerts a tumor suppressor role in lung cancer by transcriptionally downregulating cyclin D3 | ||
Cell Death Dis Identification of STAM-binding protein as a target for the treatment of gemcitabine resistance pancreatic cancer in a nutrient-poor microenvironment
| ||
Cell Rep FAK-mediated phosphorylation at Y464 regulates p85β nuclear translocation to promote tumorigenesis of ccRCC by repressing RB1 expression |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH Umut (Verified Customer) (08-30-2023) | It works very well, even though it is diluted a lot, i.e., 1:2000. The total protein concentration of my samples were around 2 ug/ml before adding 1:4 laemmli buffer and denature, and it is able to capture the target at low exposure times.
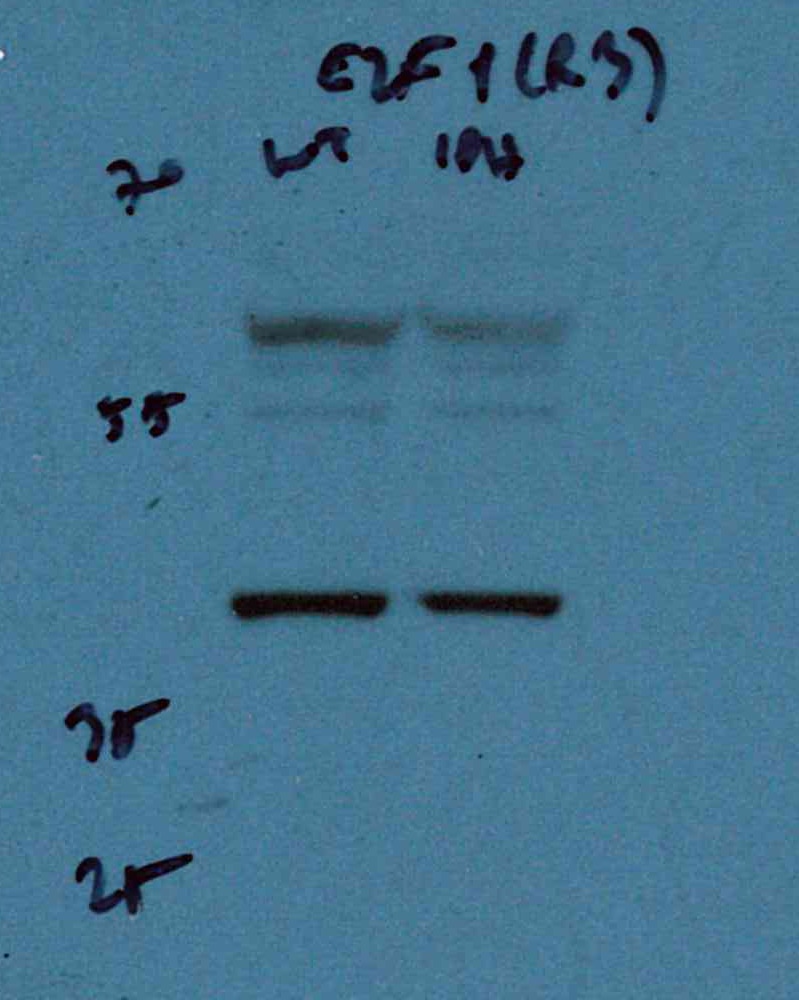 |
FH Juliane (Verified Customer) (05-11-2023) | The antibody worked fine, but only under using ECL sensitve reagent and high exposure when imaging.
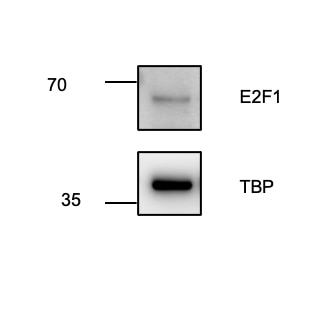 |