Tested Applications
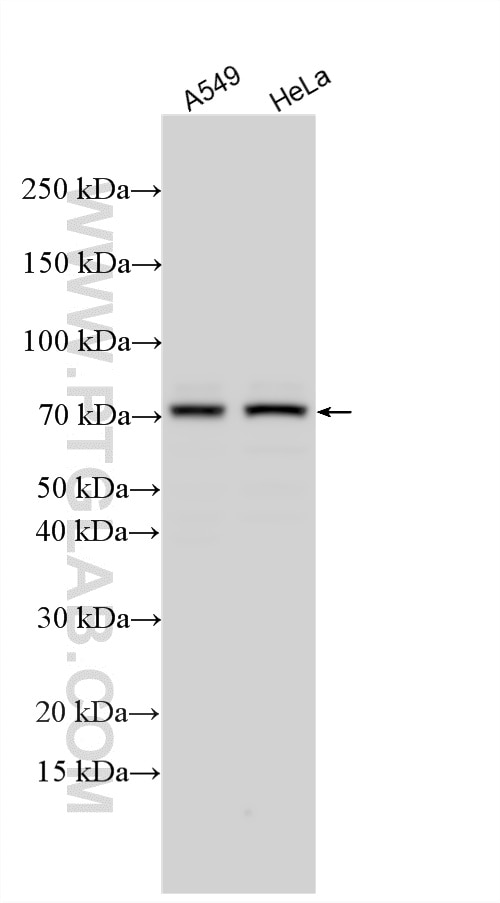
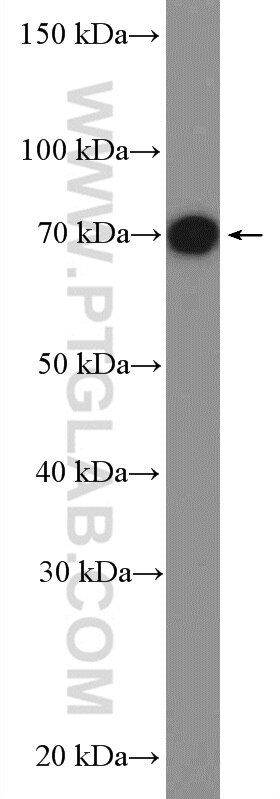
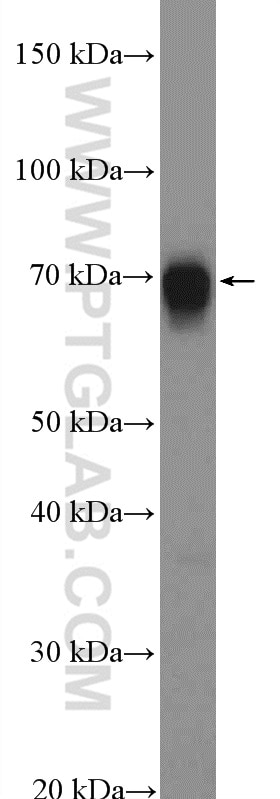
| Positive WB detected in | A549 cells, mouse colon tissue, NIH/3T3 cells, HeLa cells |
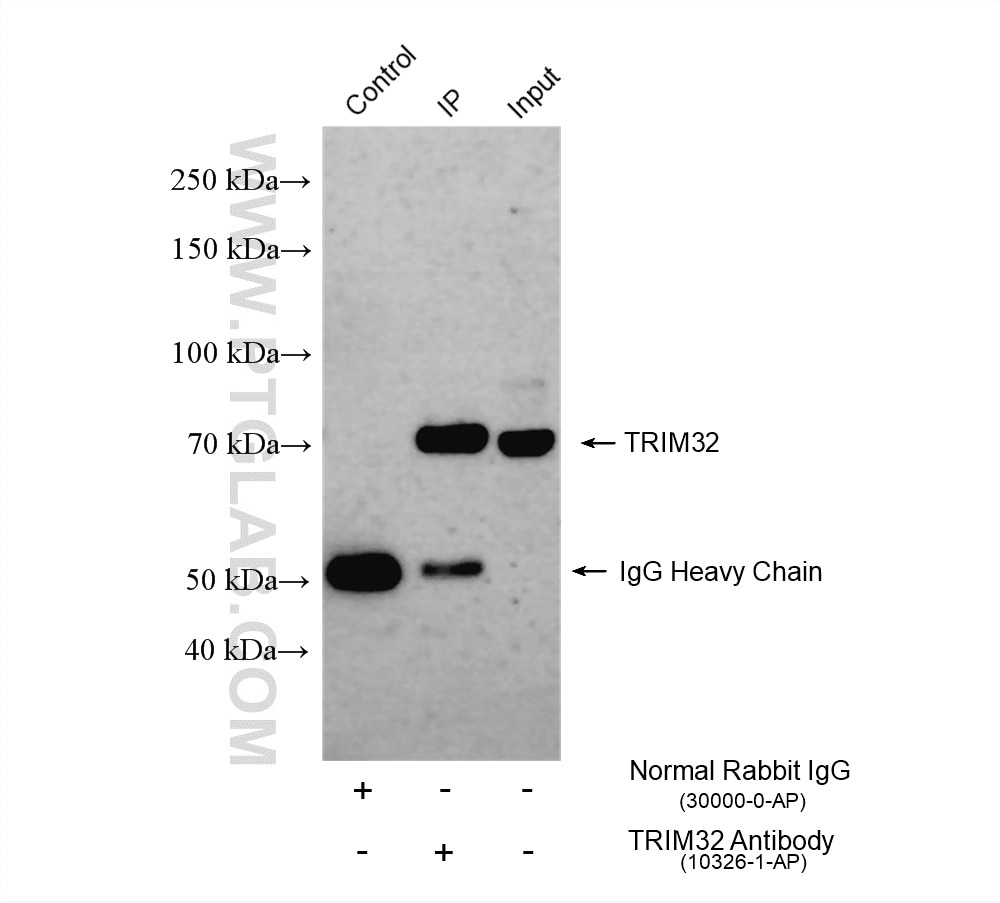
| Positive IP detected in | U-251 cells |
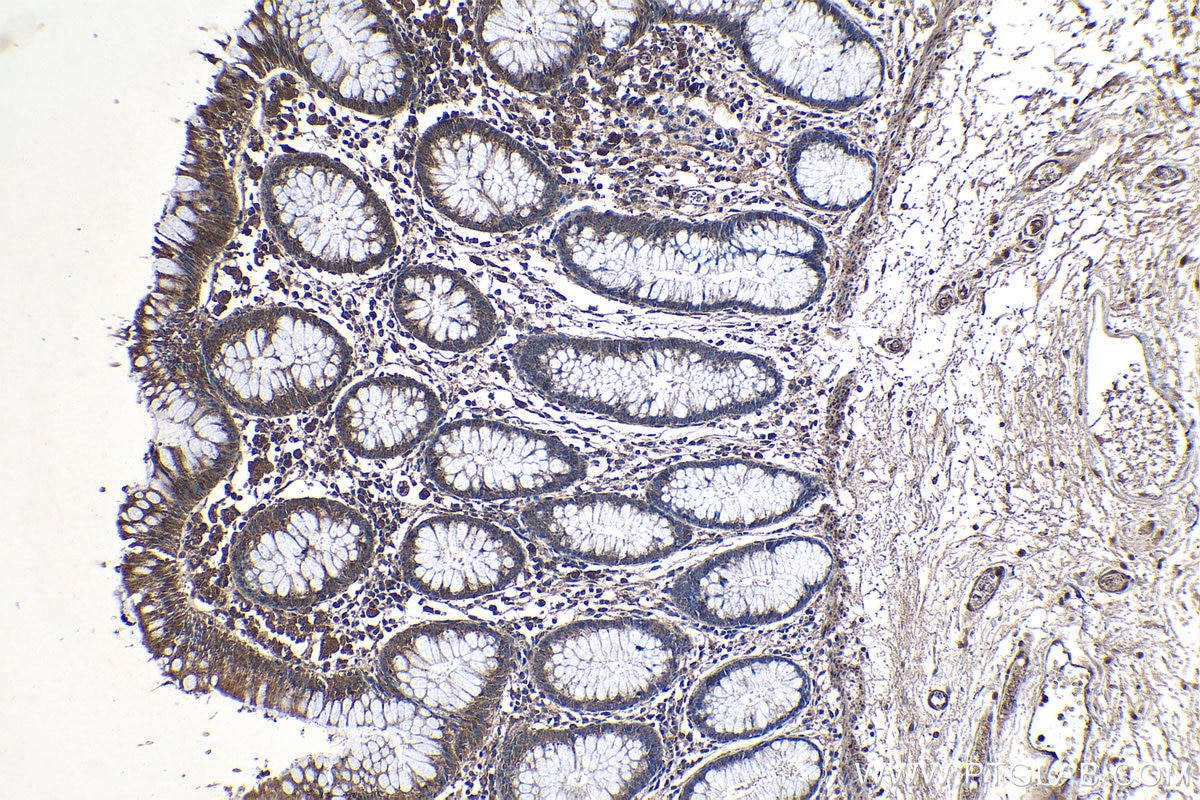
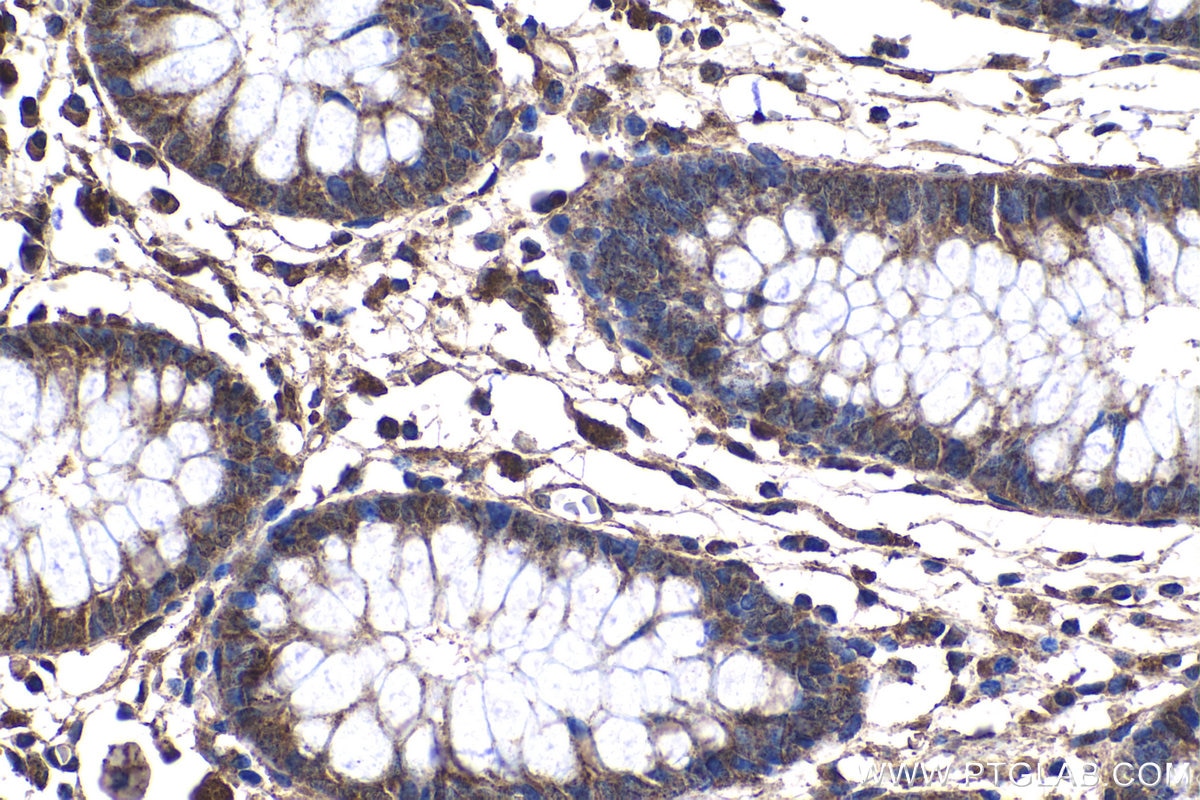
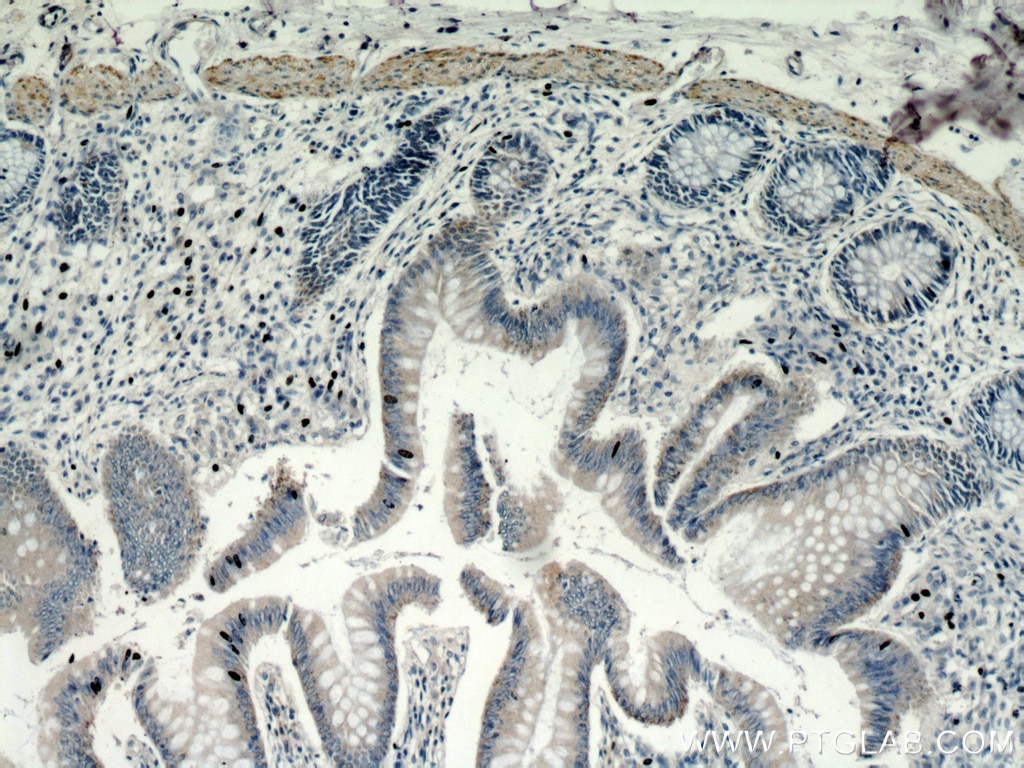
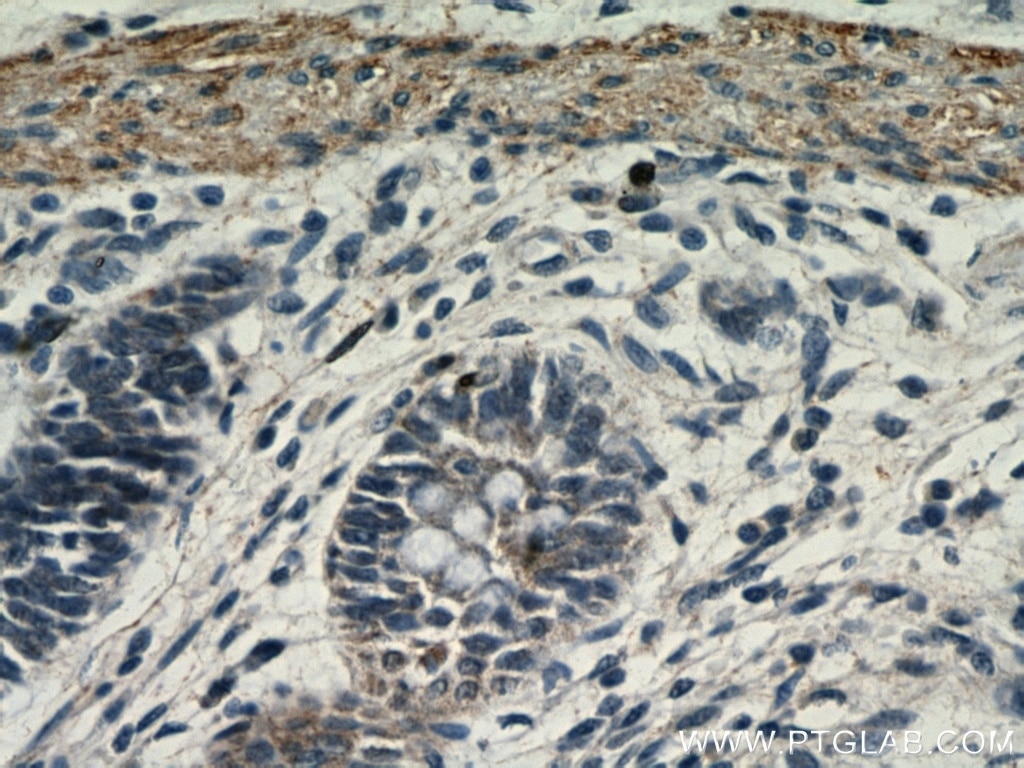
| Positive IHC detected in | human colon tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
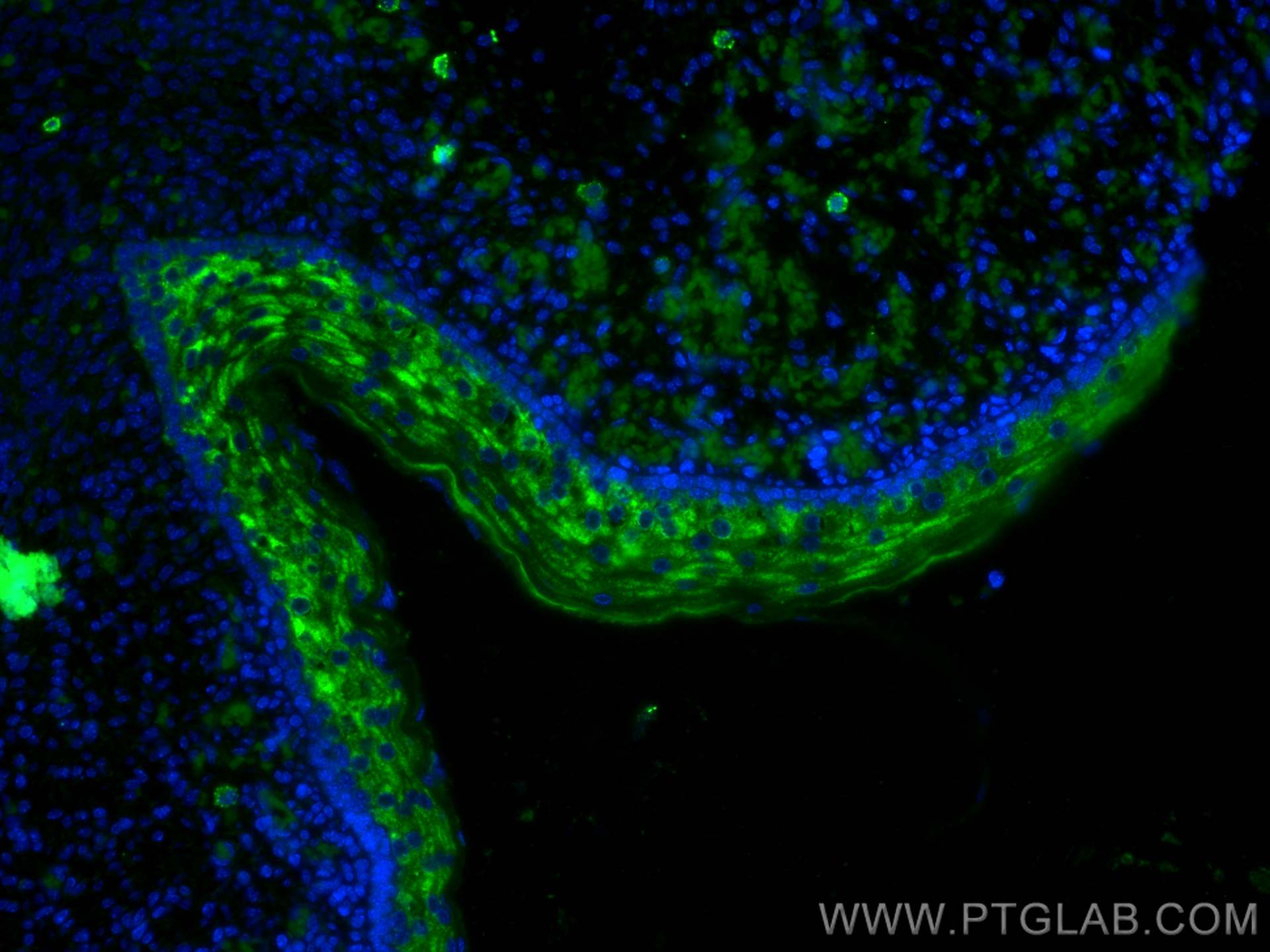
| Positive IF-P detected in | mouse embryo tissue |
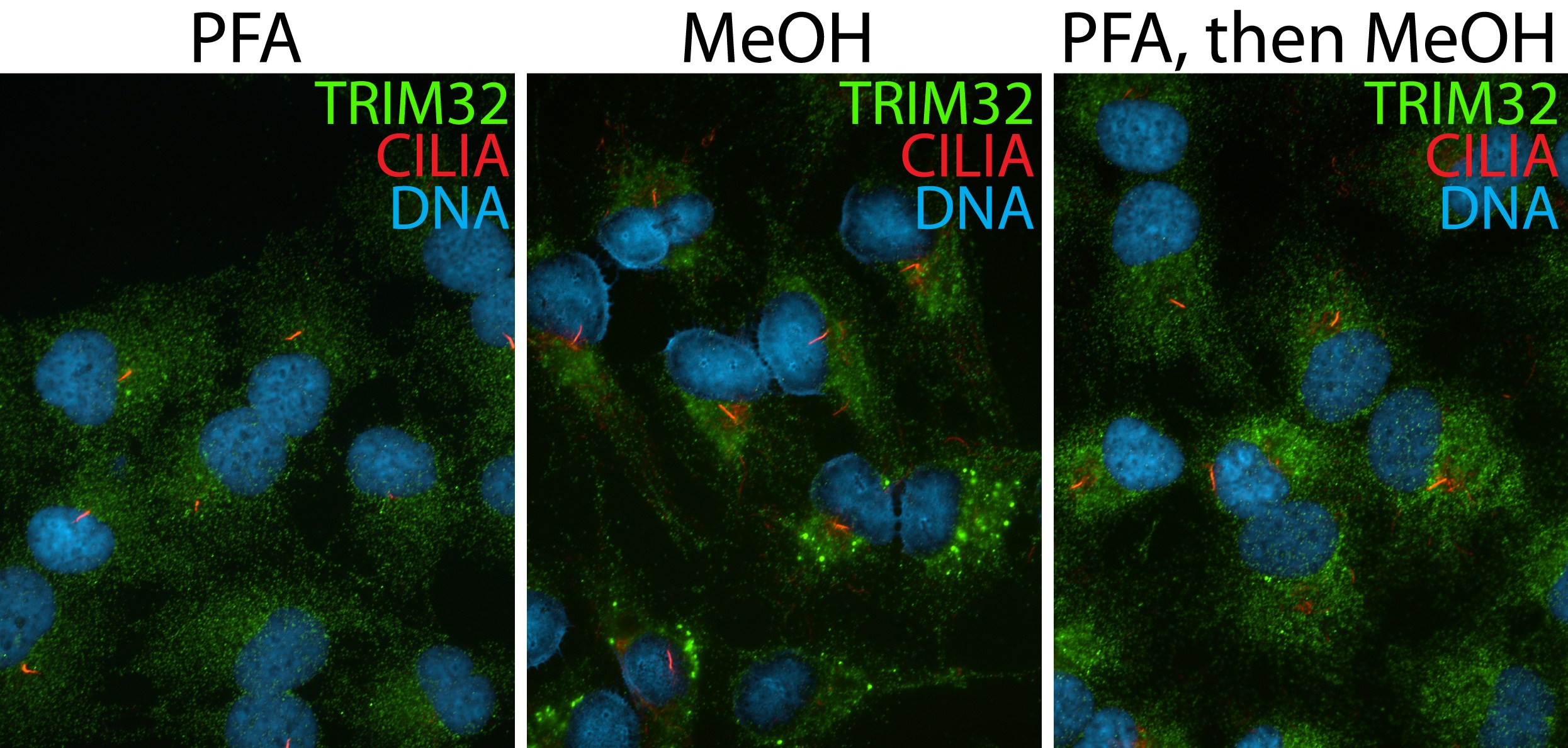
| Positive IF/ICC detected in | hTERT-RPE1 cells |
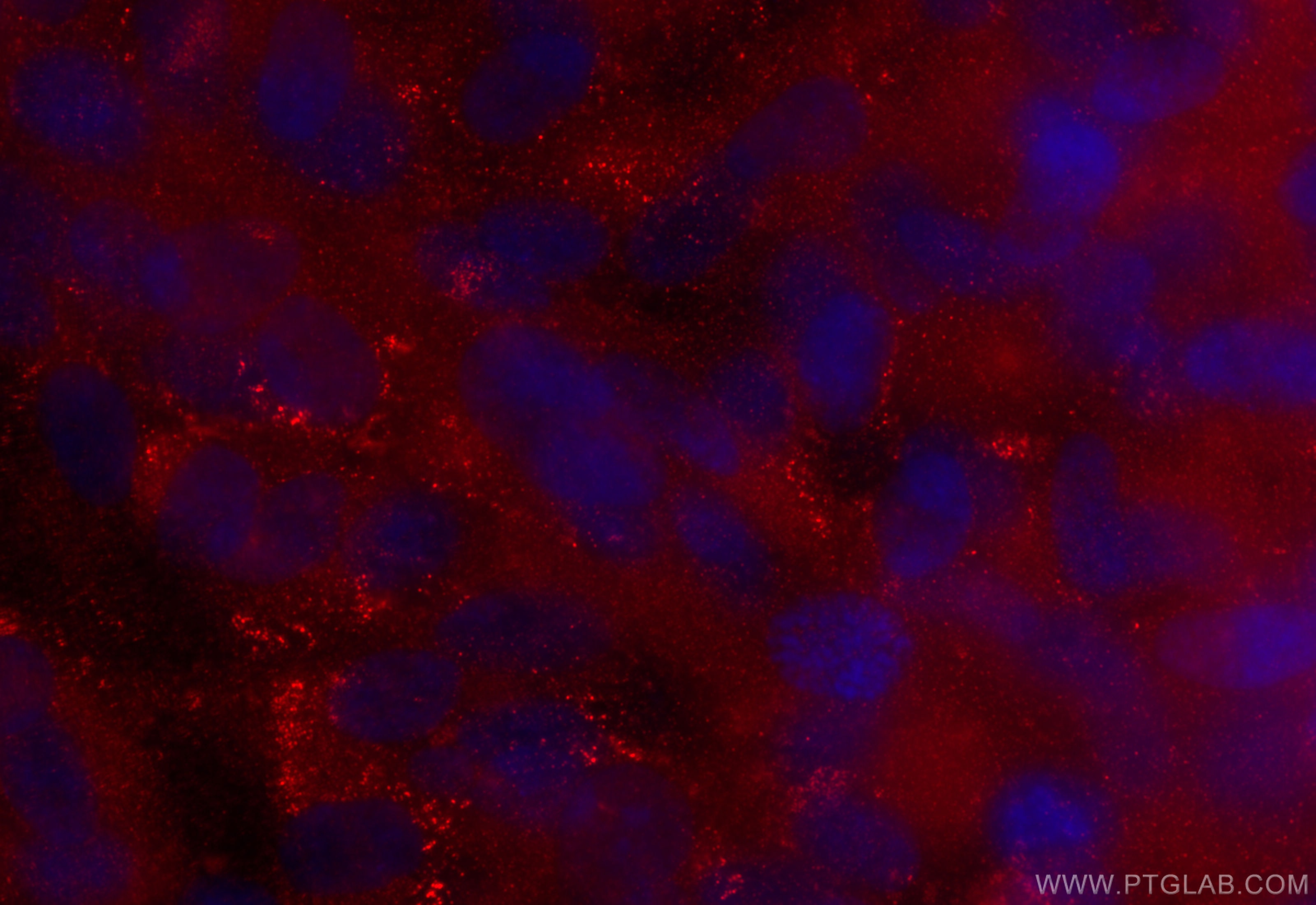
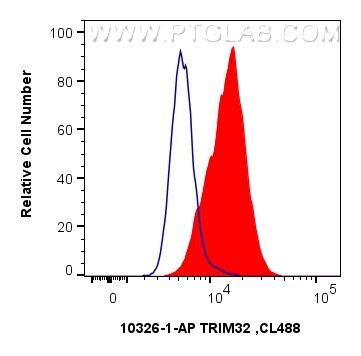
| Positive FC (Intra) detected in | A431 cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:1000-1:8000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Immunohistochemistry (IHC) | IHC : 1:50-1:500 |
| Immunofluorescence (IF)-P | IF-P : 1:50-1:500 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:100-1:400 |
| Flow Cytometry (FC) (INTRA) | FC (INTRA) : 0.40 ug per 10^6 cells in a 100 µl suspension |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 11 publications below |
| WB | See 15 publications below |
| IHC | See 4 publications below |
| IF | See 5 publications below |
| IP | See 1 publications below |
Product Information
10326-1-AP targets TRIM32 in WB, IHC, IF/ICC, IF-P, FC (Intra), IP, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag0401 Product name: Recombinant human TRIM32 protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 111-430 aa of BC003154 Sequence: FCRSCGLVLCEPCREADHQPPGHCTLPVKEAAEERRRDFGEKLTRLRELMGELQRRKAALEGVSKDLQARYKAVLQEYGHEERRVQDELARSRKFFTGSLAEVEKSNSQVVEEQSYLLNIAEVQAVSRCDYFLAKIKQADVALLEETADEEEPELTASLPRELTLQDVELLKVGHVGPLQIGQAVKKPRTVNVEDSWAMEATASAASTSVTFREMDMSPEEVVASPRASPAKQRGPEAASNIQQCLFLKKMGAKGSTPGMFNLPVSLYVTSQGEVLVADRGNYRIQVFTRKGFLKEIRRSPSGIDSFVLSFLGADLPNLT Predict reactive species |
| Full Name | tripartite motif-containing 32 |
| Calculated Molecular Weight | 72 kDa |
| Observed Molecular Weight | 70-72 kDa |
| GenBank Accession Number | BC003154 |
| Gene Symbol | TRIM32 |
| Gene ID (NCBI) | 22954 |
| RRID | AB_2208005 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q13049 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
TRIM32(Tripartite motif-containing protein 32) is a widely expressed ubiquitin ligase that is localized to the Z-line in skeletal muscle(PMID:19349376). It promotes PIAS4 ubiquitination and degradation, and by controlling PIAS4 stability, regulates UVB-induced keratinocyte apoptosis through induction of NFkappaB(PMID: 16816390). Western blot analysis detected TRIM32 at an apparent molecular mass of 70-72 kDa.
Protocols
| Product Specific Protocols | |
|---|---|
| FC protocol for TRIM32 antibody 10326-1-AP | Download protocol |
| IF protocol for TRIM32 antibody 10326-1-AP | Download protocol |
| IHC protocol for TRIM32 antibody 10326-1-AP | Download protocol |
| IP protocol for TRIM32 antibody 10326-1-AP | Download protocol |
| WB protocol for TRIM32 antibody 10326-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cell Death Differ TRIM32 promotes anoikis resistance and metastasis in NSCLC by degrading CHEK2 to enhance IL-6 secretion
| ||
Sci Adv An EMT-primary cilium-GLIS2 signaling axis regulates mammogenesis and claudin-low breast tumorigenesis. | ||
J Am Soc Nephrol Proteome Analysis of Isolated Podocytes Reveals Stress Responses in Glomerular Sclerosis. | ||
Hum Mol Genet Deficiency of the E3 ubiquitin ligase TRIM32 in mice leads to a myopathy with a neurogenic component.
| ||
PLoS Pathog TRIM32 inhibits Venezuelan equine encephalitis virus infection by targeting a late step in viral entry
| ||
FASEB J Ubiquitin ligase TRIM32 promotes dendrite arborization by mediating degradation of the epigenetic factor CDYL.
|
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH Ashraf (Verified Customer) (08-22-2024) | We have so far struggled with detecting the knockdown/knockout of Trim32 in our experimental settings despite testing several antibodies. We could very clearly see downregulation or absence of Trim32 as expected at transcript level but failed to detect the same at protein level. However, use of Trim32 Ab from Proteintech has unbelievably solved this issue! Highly highly recommended *****
|