Tested Applications
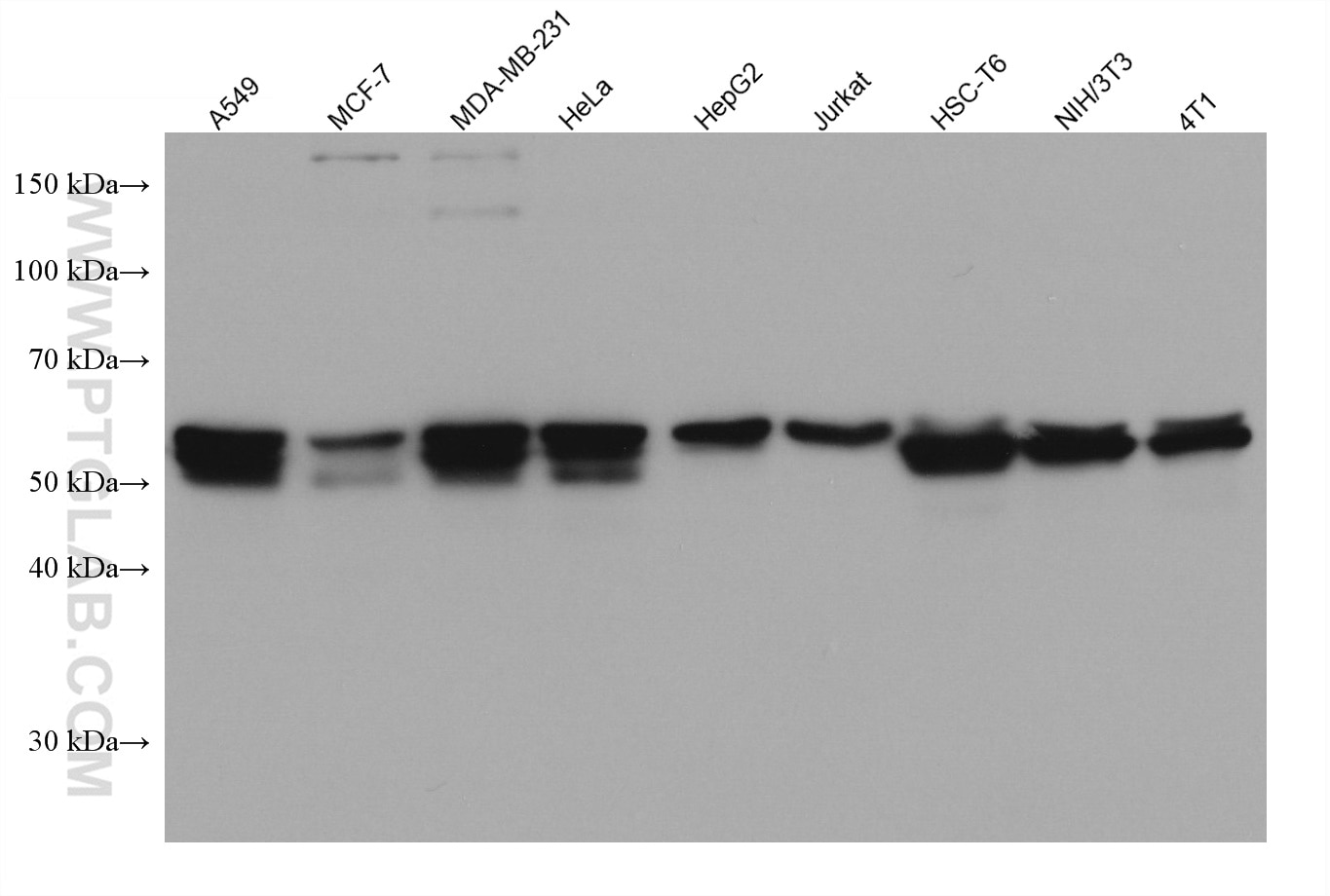
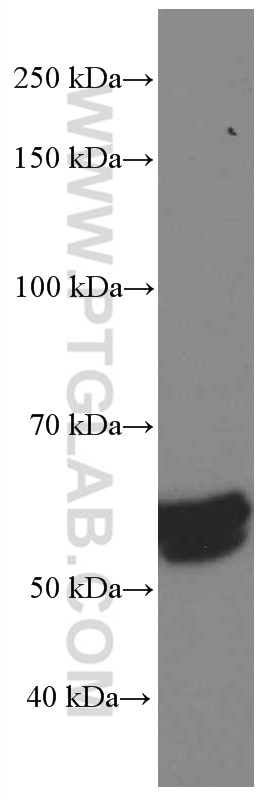
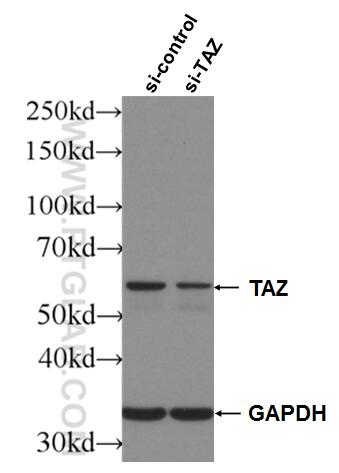
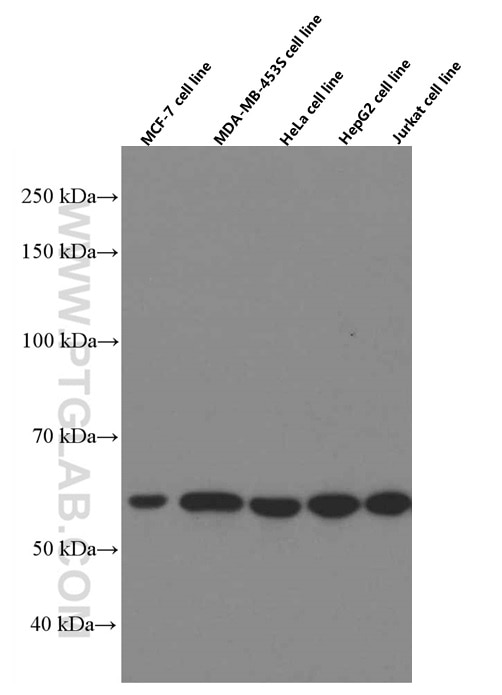
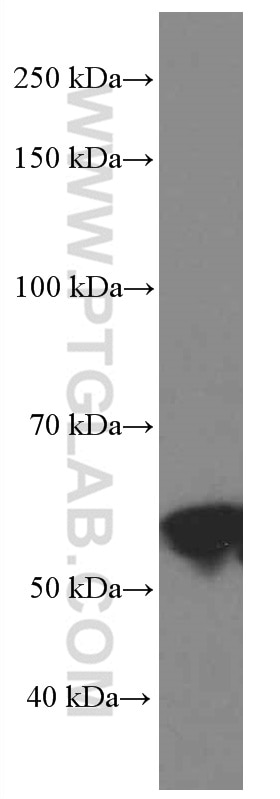
| Positive WB detected in | A549 cells, HepG2 cells, HSC-T6 cells, MCF-7 cells, NIH/3T3 cells, MDA-MB-231 cells, HeLa cells, Jurkat cells, 4T1 cells, MDA-MB-453S cells |
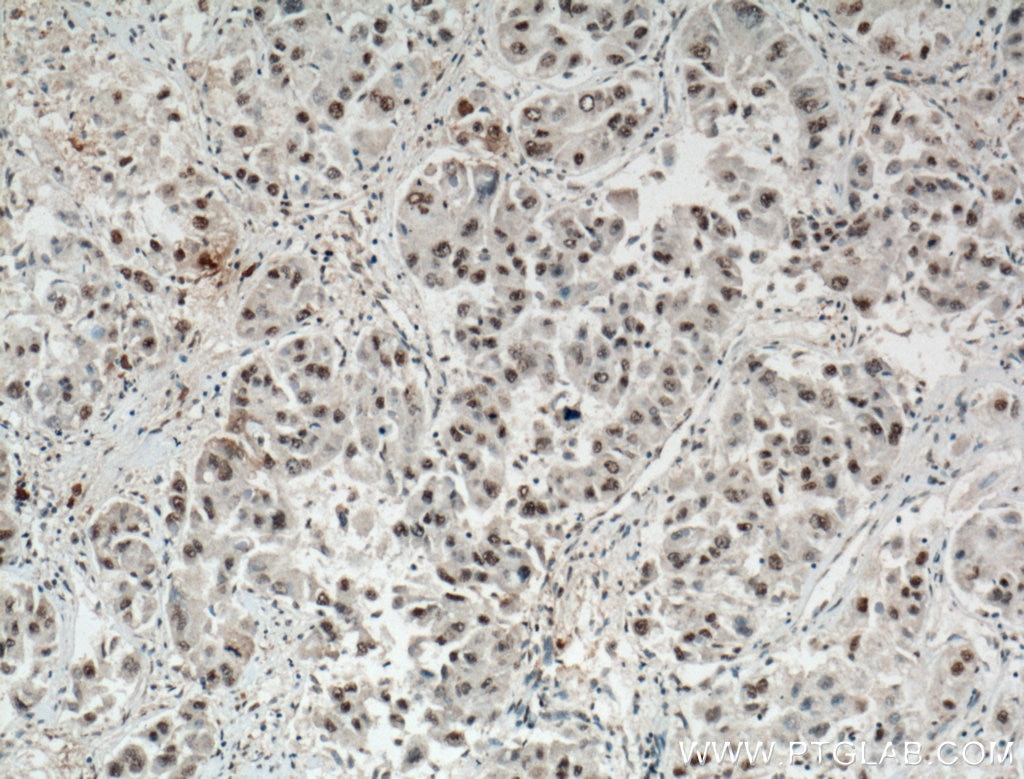
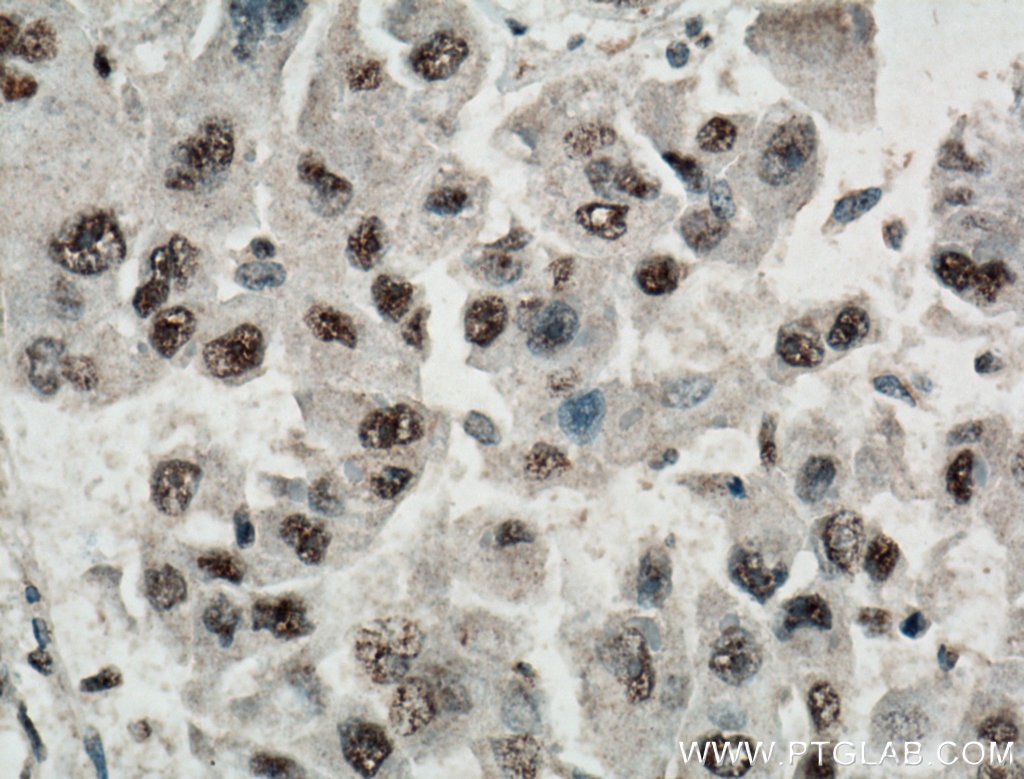
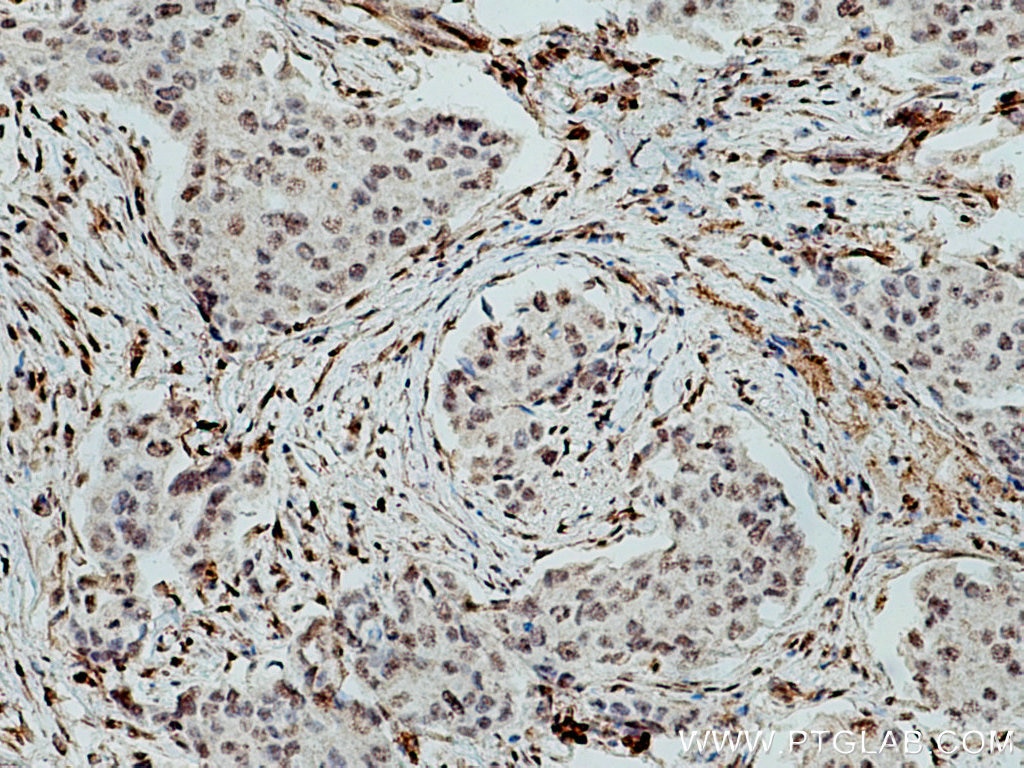
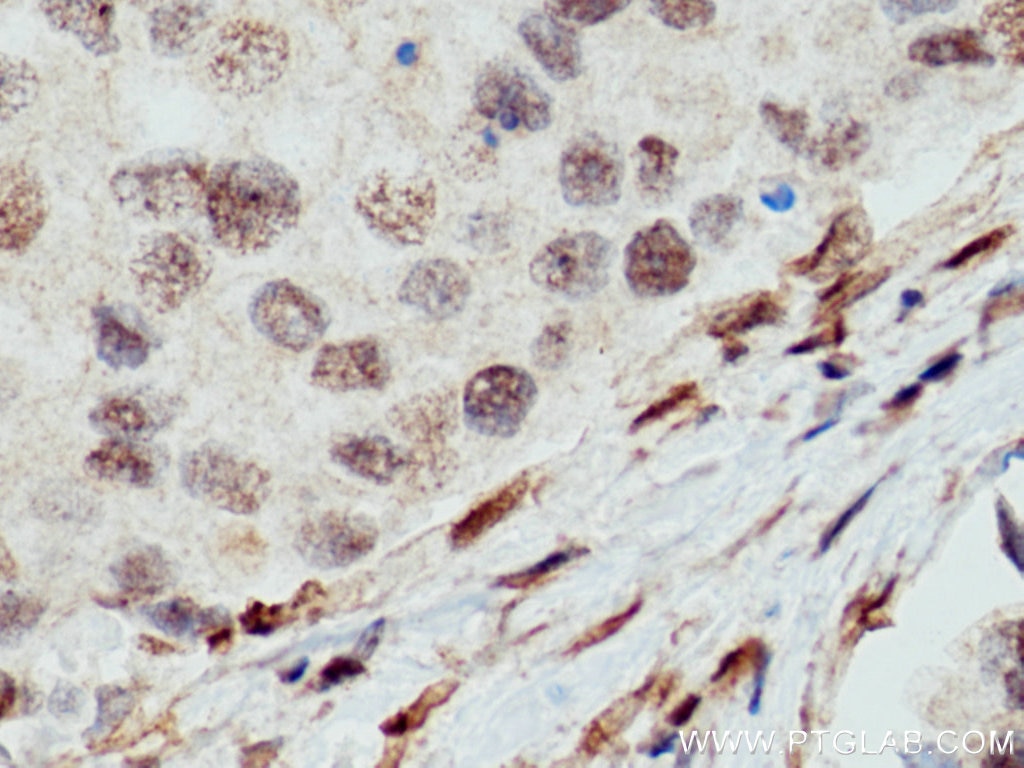
| Positive IHC detected in | human liver cancer tissue, human breast cancer tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
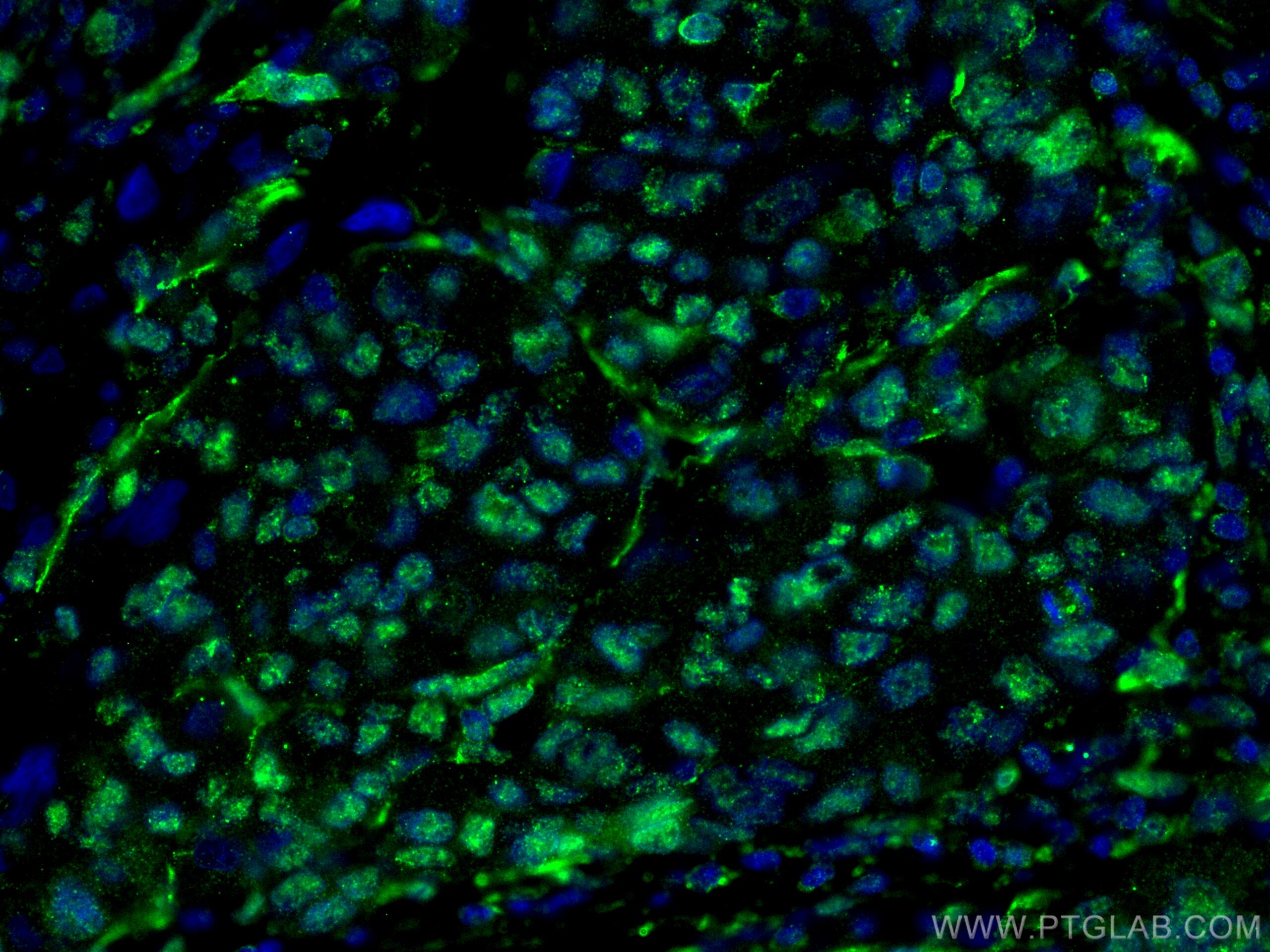
| Positive IF-P detected in | human liver cancer tissue |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:5000-1:50000 |
| Immunohistochemistry (IHC) | IHC : 1:300-1:1200 |
| Immunofluorescence (IF)-P | IF-P : 1:200-1:800 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 1 publications below |
| WB | See 15 publications below |
| IHC | See 3 publications below |
| IF | See 9 publications below |
Product Information
66500-1-Ig targets WWTR1/TAZ in WB, IHC, IF-P, ELISA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Mouse / IgG1 |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag13330 Product name: Recombinant human WWTR1,TAZ protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 1-400 aa of BC014052 Sequence: MNPASAPPPLPPPGQQVIHVTQDLDTDLEALFNSVMNPKPSSWRKKILPESFFKEPDSGSHSRQSSTDSSGGHPGPRLAGGAQHVRSHSSPASLQLGTGAGAAGSPAQQHAHLRQQSYDVTDELPLPPGWEMTFTATGQRYFLNHIEKITTWQDPRKAMNQPLNHMNLHPAVSSTPVPQRSMAVSQPNLVMNHQHQQQMAPSTLSQQNHPTQNPPAGLMSMPNALTTQQQQQQKLRLQRIQMERERIRMRQEELMRQEAALCRQLPMEAETLAPVQAAVNPPTMTPDMRSITNNSSDPFLNGGPYHSREQSTDSGLGLGCYSVPTTPEDFLSNVDEMDTGENAGQTPMNINPQQTRFPDFLDCLPGTNVDLGTLESEDLIPLFNDVESALNKSEPFLTWL Predict reactive species |
| Full Name | WW domain containing transcription regulator 1 |
| Calculated Molecular Weight | 44 kDa |
| Observed Molecular Weight | 55 kDa |
| GenBank Accession Number | BC014052 |
| Gene Symbol | WWTR1 |
| Gene ID (NCBI) | 25937 |
| RRID | AB_2881864 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein G purification |
| UNIPROT ID | Q9GZV5 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
TAZ, also referred as WWTR1, is a transcriptional co-activator with a PDZ-binding motif that is regulated by its interaction with 14-3-3 protein. TAZ is expressed in many primary tumors, such as breast cancer, thyroid carcinoma, colorectal cancer, and glioma. TAZ has been reported to be one of the nuclear effectors of Hippo-related pathways that regulate organ size control, cancer stem cell (CSC) properties, and epithelial-mesenchymal transition (EMT). TAZ has been defined for its role in the nucleus, where it functions directly as a transcriptional regulator by interacting with several nuclear factors as Runx2/Cbfa1, NHERF-2, TEF-1, TBX5, PAX3, PAX8 and TTF-1.
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for WWTR1/TAZ antibody 66500-1-Ig | Download protocol |
| IHC protocol for WWTR1/TAZ antibody 66500-1-Ig | Download protocol |
| WB protocol for WWTR1/TAZ antibody 66500-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cell Death Differ Targeting USP10 induces degradation of oncogenic ANLN in esophageal squamous cell carcinoma | ||
Nat Commun LPA signaling acts as a cell-extrinsic mechanism to initiate cilia disassembly and promote neurogenesis.
| ||
Aging Cell Inhibition of Hippo Signaling Through Ablation of Lats1 and Lats2 Protects Against Cognitive Decline in 5xFAD Mice via Increasing Neuronal Resilience Against Ferroptosis | ||
Cell Signal RhoA/ROCK-TAZ Axis regulates bone formation within calvarial trans-sutural distraction osteogenesis | ||
Front Pharmacol TAZ Induces Migration of Microglia and Promotes Neurological Recovery After Spinal Cord Injury. | ||
Am J Transl Res VASN promotes YAP/TAZ and EMT pathway in thyroid carcinogenesis in vitro. |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
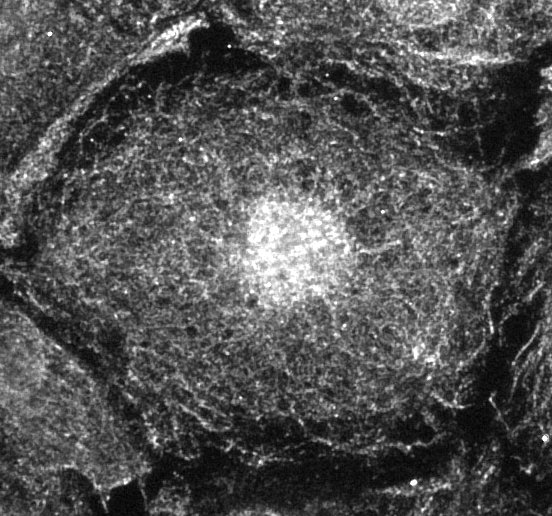
FH Rumeyza (Verified Customer) (03-26-2026) | the immunostaining worked well.
 |