mNeonGreen vs GFP
mNeonGreen: A green fluorescent protein that is not a GFP variant
mNeon-Green is a new bright monomeric yellow-green fluorescent protein. It has already been frequently used for mainly microscopic applications in both wide-field microscopy and super-resolution microscopy. What is mNeonGreen all about?
mNeonGreen vs. GFP – What are the differences?
About GFP
Green fluorescent protein (GFP) has been originally isolated from jellyfish Aequorea Victoria. Frequently, it has been genetically modified to generate a large number of GFP variants with special spectral or functional properties. The most common variant EGFP has an excitation peak at 488 nm and an emission peak at 509 nm. The filter settings of most microscopes are optimized for those wavelengths. Learn more about GFP here.
Evolutionary distance of mNeonGreen to jellyfish GFP
mNeonGreen is also a protein that is fluorescent in the green range, but it is not a variant of GFP. Instead, mNeonGreen is derived from the lancelet Branchiostoma lanceolatum multimeric yellow fluorescence protein. Hence, mNeonGreen is evolutionarily distant from jellyfish-derived GFP. At sequence level, mNeonGreen shares just 20-25% sequence identity with common EGFP:
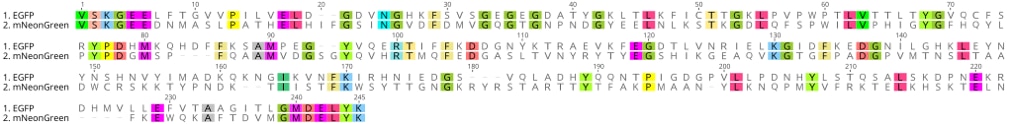
The mNeonGreen DNA sequence can be found here: https://www.ncbi.nlm.nih.gov/nuccore/KC295282.1 or https://www.ebi.ac.uk/ena/data/view/KC295282
Excitation & Emission wavelengths and brightness of mNeonGreen
mNeonGreen has an excitation maximum at 506 nm and an emission maximum at 517 nm (Shaner et al., 2013). Therefore, mNeonGreen is compatible with most GFP filter sets. mNeonGreen is also 3 times brighter than GFP when using GFP filters; optimization of filters may increase its brightness further.
mNeonGreen seems to be more stable and less sensitive to laser induced bleaching than EGFP. Therefore, mNeonGreen is suitable for confocal and super resolution microscopy, especially when fusion proteins are investigated, which are expressed at low levels.
Size of mNeonGreen
mNeonGreen consists of 237 amino acids, which translates into 26.6 kDa molecular weight. This size, but not sequence, is very similar to EGFP that comprises 239 amino acids and is 26.9 kDa.
Applications of GFP and mNeonGreen
Since its introduction as a molecular biology tool in the early 1990s, GFP and its variants have been extensively used for a multitude of applications such as
- Immunofluorescence (wide field, confocal & super resolution microscopy)
- Immunoprecipitation & Co-immunoprecipitation
- Western blotting
- Mass spectrometry
- Enzyme activity measurements
- RIP/ ChIP analysis
- Reporter assays
In order to deal with this variety of experimental options, applications optimized GFP derivatives have been generated: these comprise modified spectral properties, e.g. large stokes shift, split variants, pH sensitive, fast folding, redox sensitive, photoactivatable, etc. variants. In addition, various tools for GFP have been developed to support each application.
So far, mNeonGreen is mainly used for expression as fusion protein in cells, tissues, or organs and analysis thereof in wide field, confocal & super resolution microscopy. mNeonGreen seems to be more stable in STED and allows to take more images. Its expression pattern is identical or very similar for strongly expressed proteins (Hostettler et al., 2017).
Research tools for mNeon Green
Due to the low sequence similarity to mNeonGreen, it is expected that affinity tools for GFP variants, i.e. antibodies, should not bind to mNeonGreen and vice versa. Accordingly, dedicated research tools have been developed for mNeonGreen:
- Immunofluorescence
- Immunoprecipitation
- mNeonGreen-Trap consists of a single-domain antibody (Nanobody or VHH) coupled to Agarose or Magnetic Agarose beads
References
N.C. Shaner et al., A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat Methods. 2013 May;10(5):407-9. doi: 10.1038/nmeth.2413. Epub 2013 Mar 24.
Hostettler et al., The Bright Fluorescent Protein mNeonGreen Facilitates Protein Expression Analysis In Vivo, G3: Genes, Genomes, Genetics February 1, 2017 vol. 7 no. 2 607-615; https://doi.org/10.1534/g3.116.038133
Related Content
Mass spec-compatible immunoprecipiation for GFP, mNeonGreen, Myc, RFP, Spot, and TurboGFP
GFP (green fluorescent protein): Properties, origin, specifications, tips
Immunoprecipitation without additional bands
GFP-Booster: GFP Nanobody for better images in immunofluorescence
GFP and RFP-Booster for better immunofluorescence imaging
Which beads should I use for my immunoprecipitation?
Support
Request a Free Nano-Trap Sample
Nano-Trap system provides fast, reliable, and effective immunoprecipitation of fusion proteins. More than 3,000 peer-reviewed articles have already been published using ChromoTek's Nano-Traps.

