Tested Applications
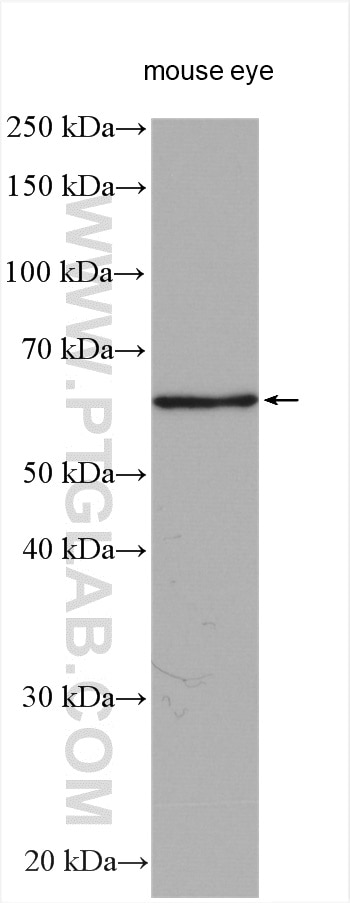
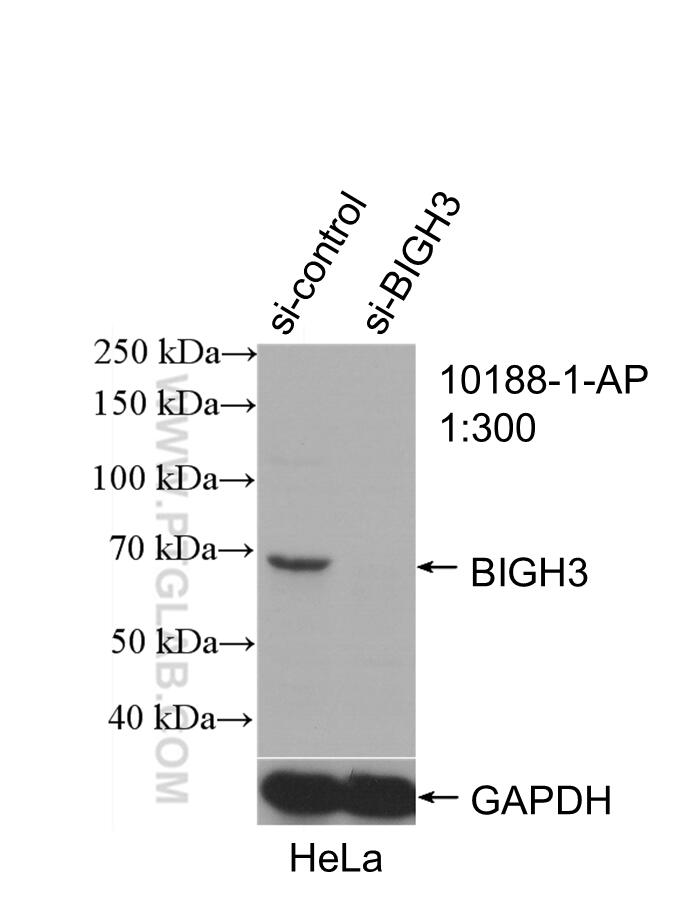
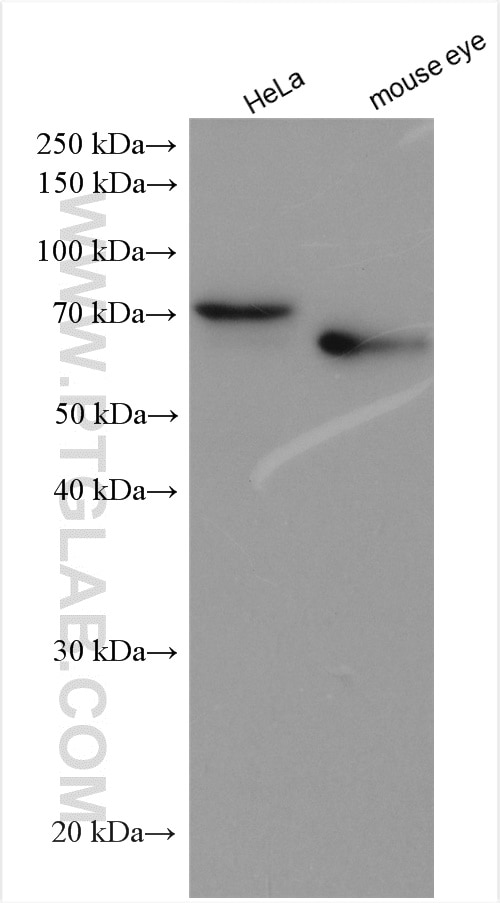
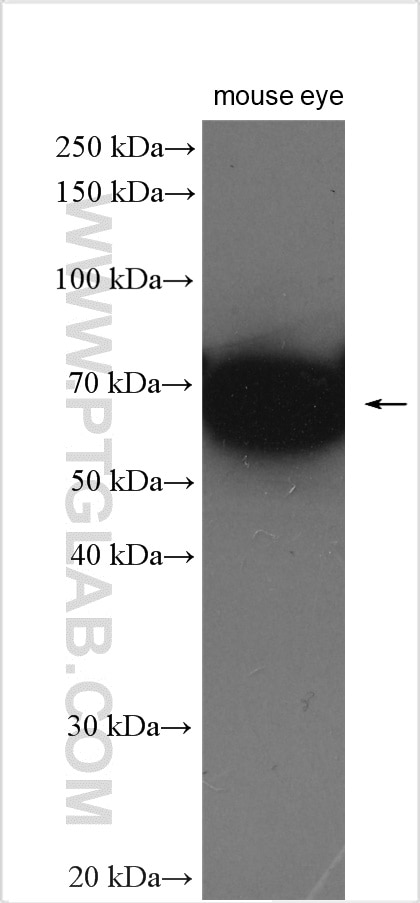
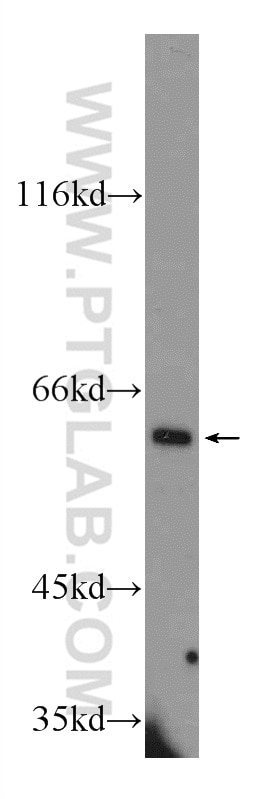
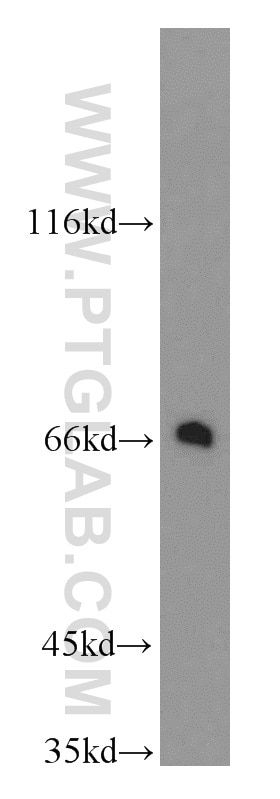
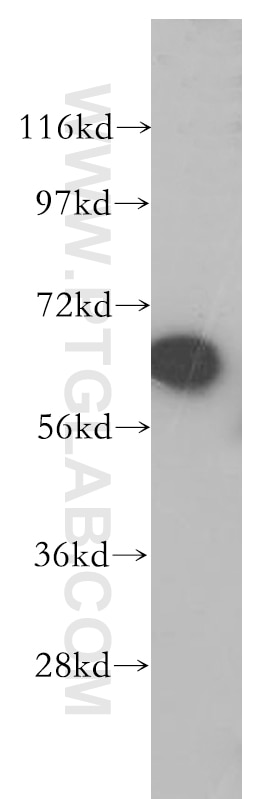
| Positive WB detected in | mouse eye tissue, mouse liver tissue, human kidney tissue, Y79 cells, HeLa cells |
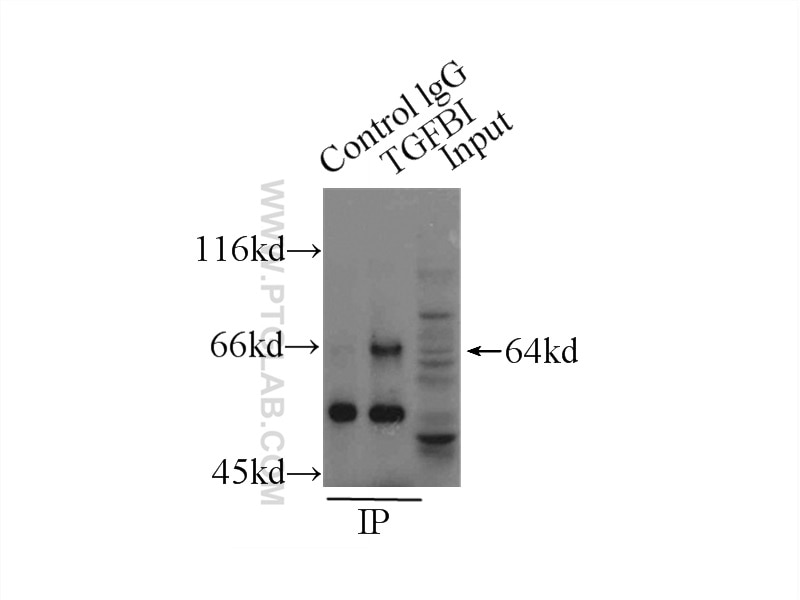
| Positive IP detected in | HeLa cells |
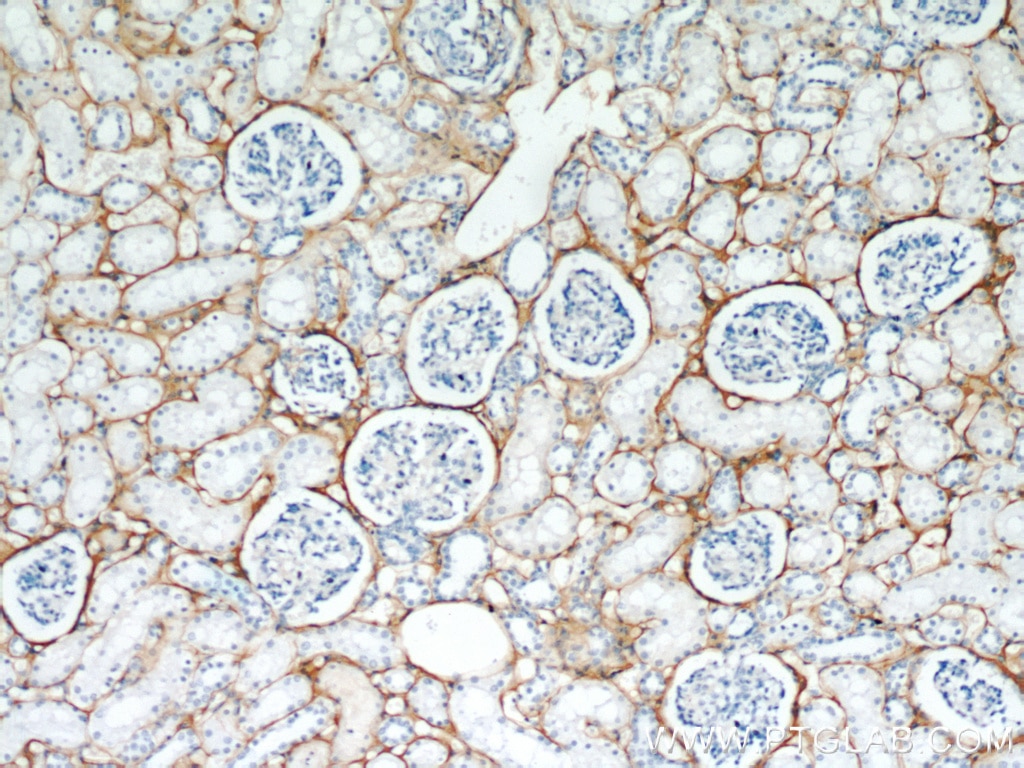
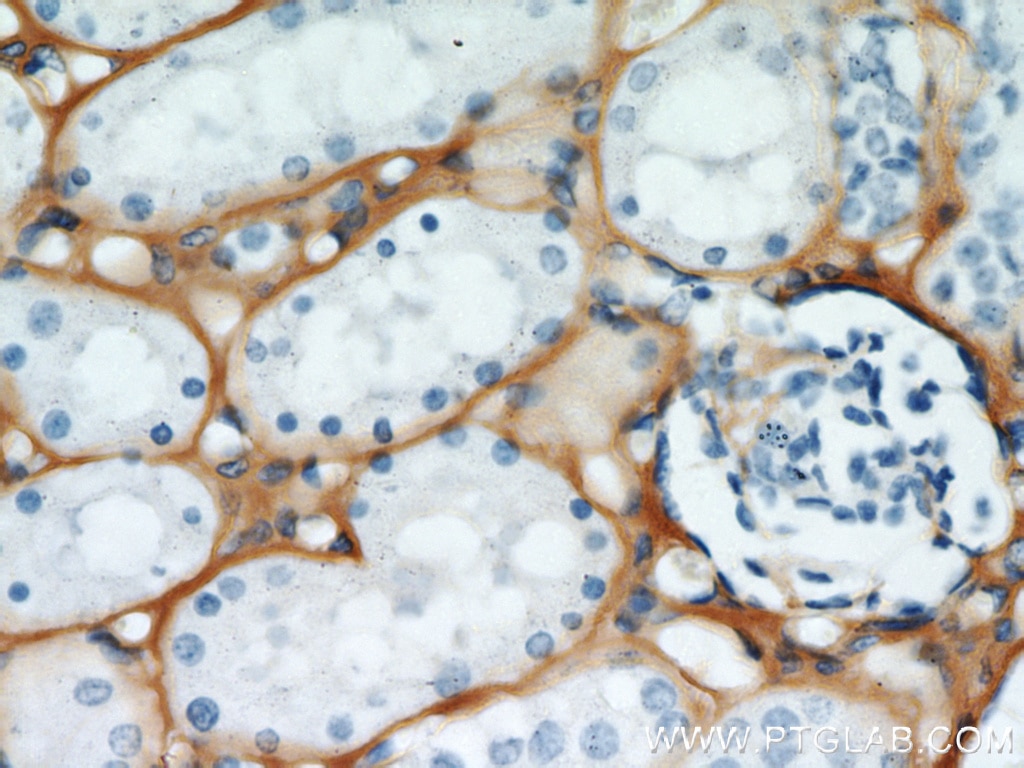
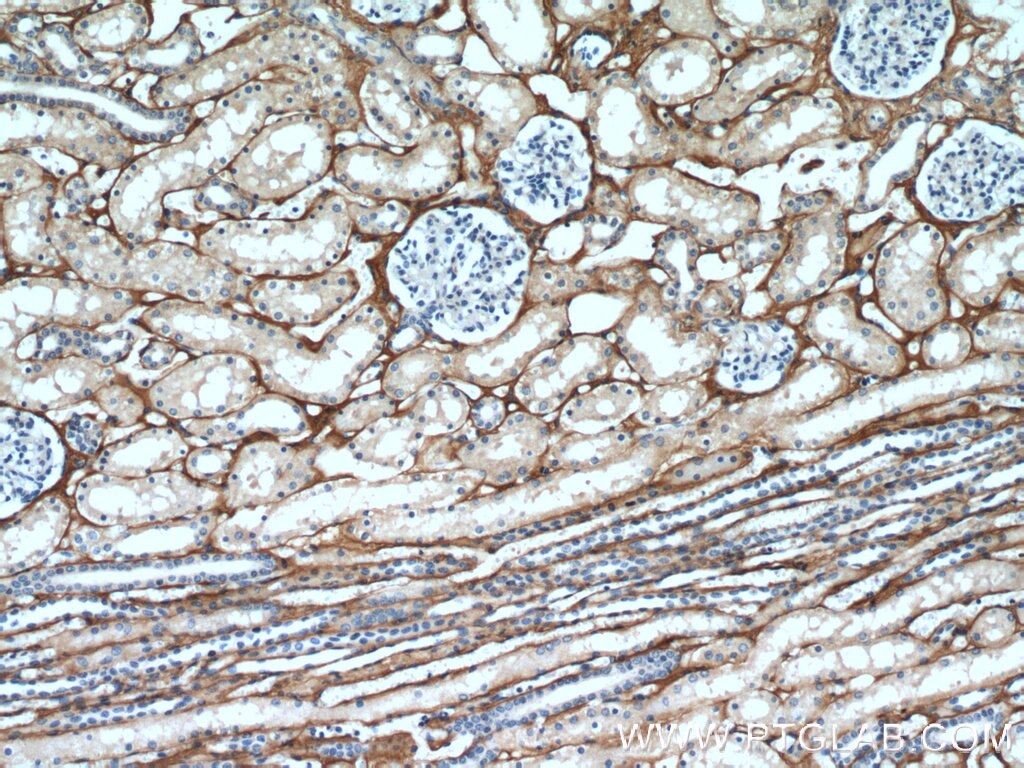
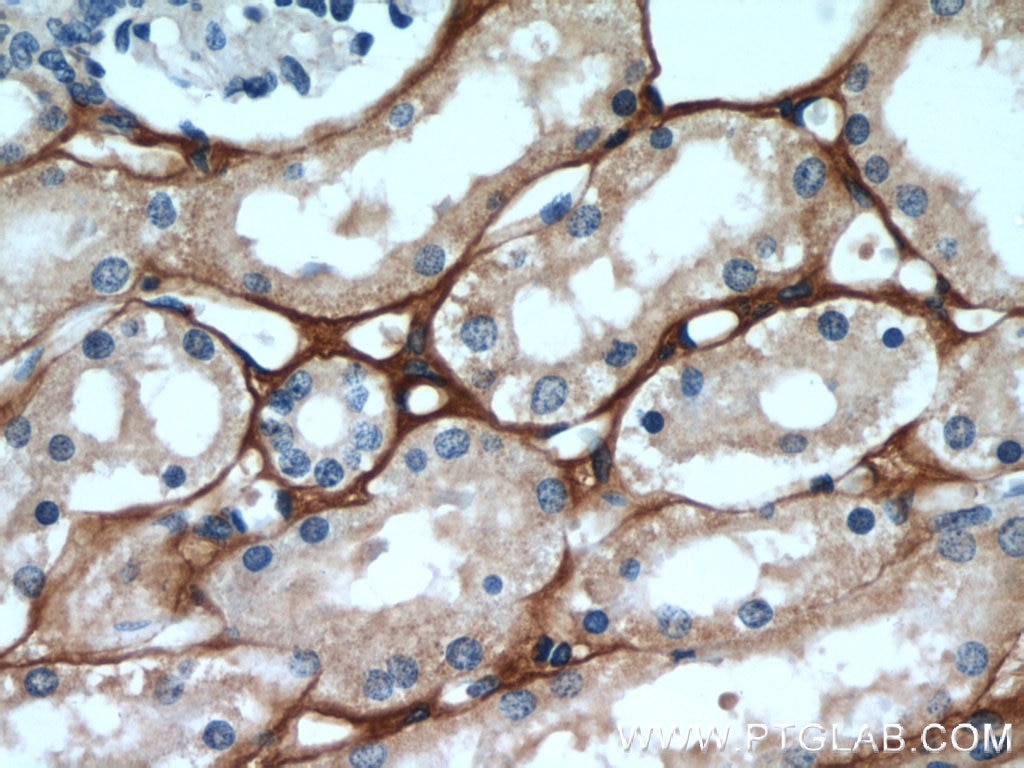
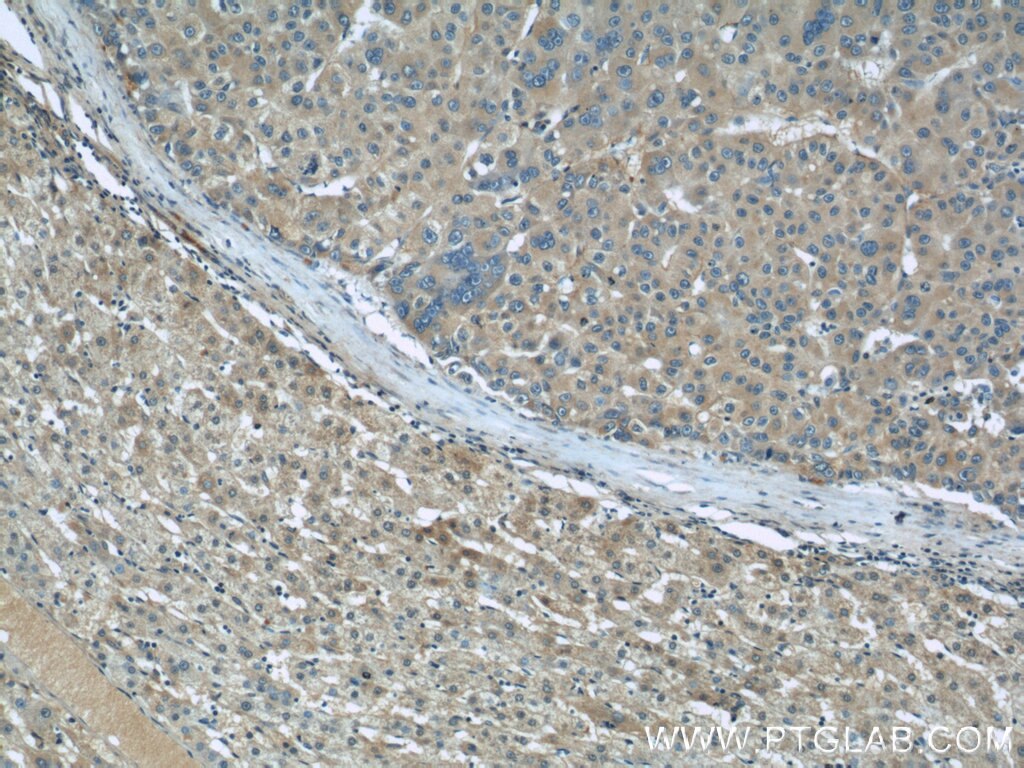
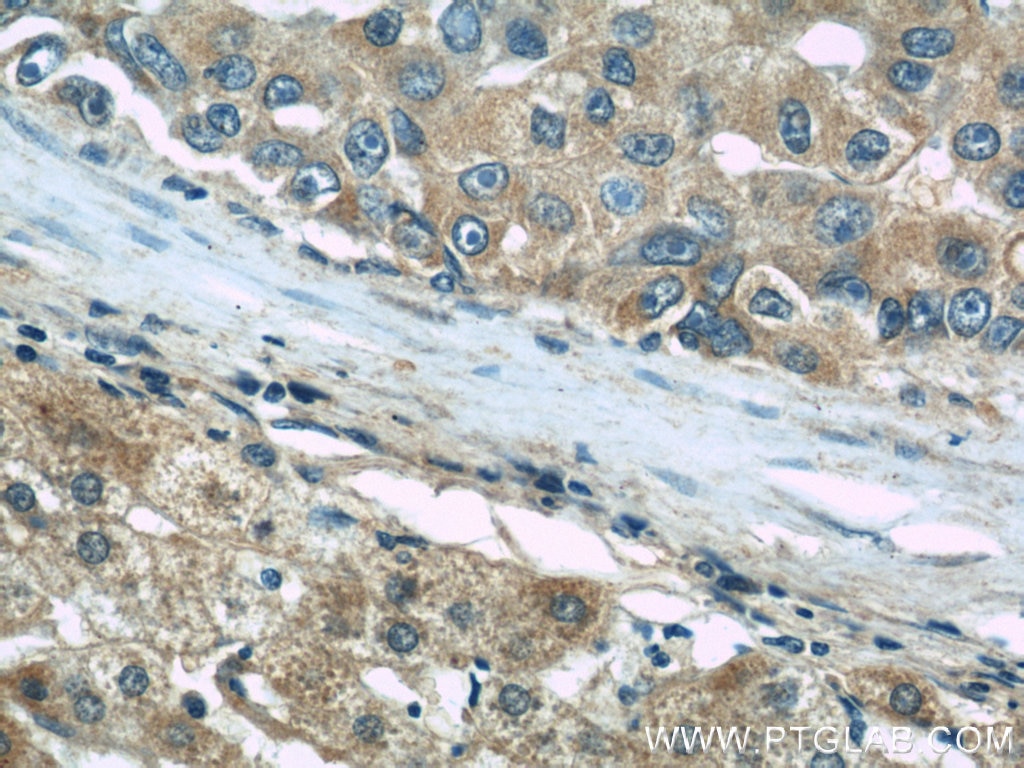
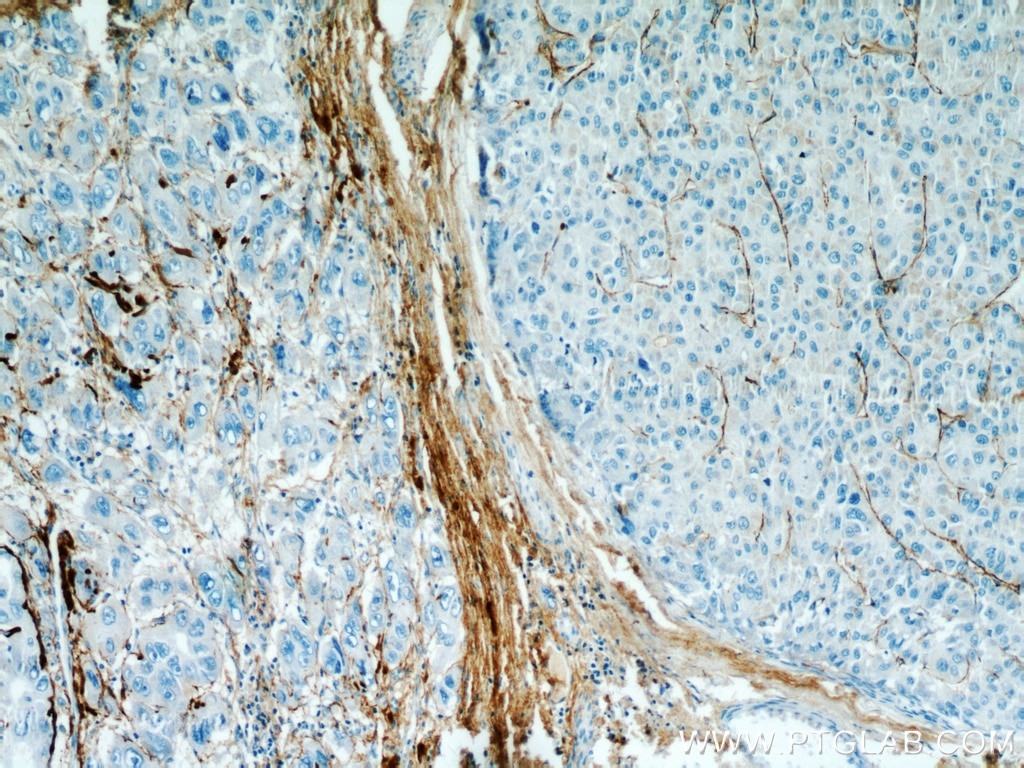
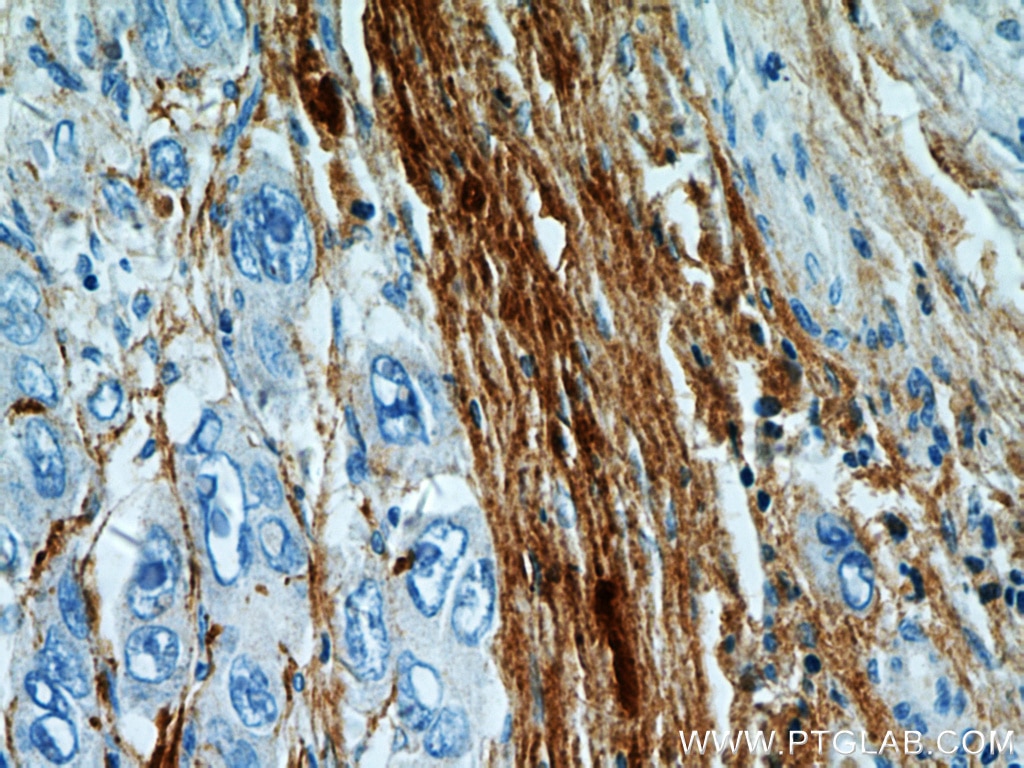
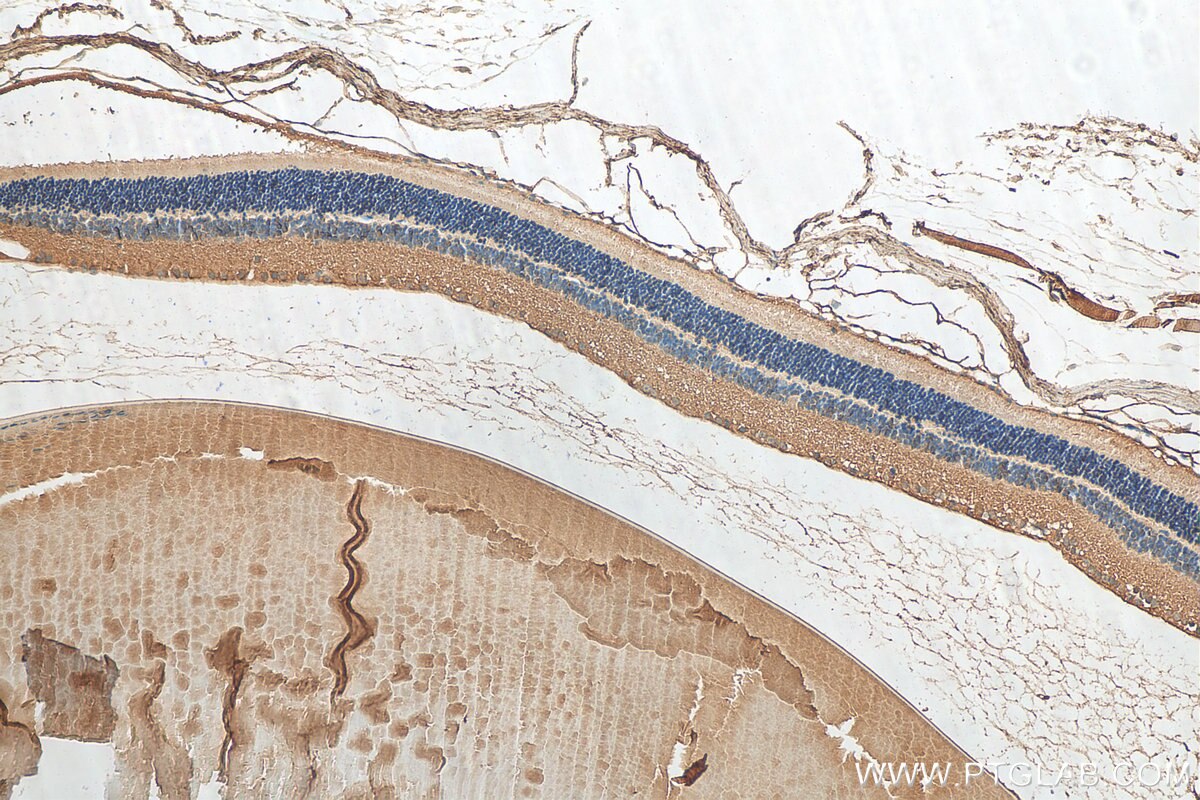
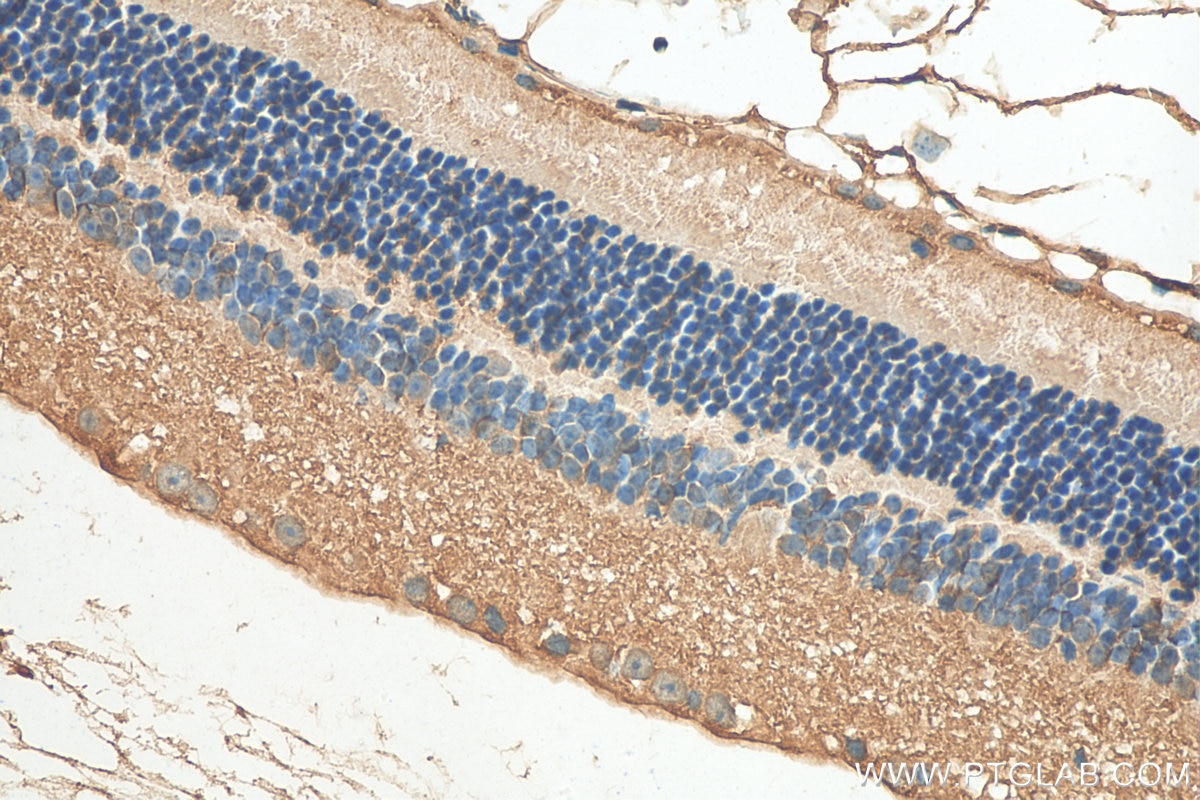
| Positive IHC detected in | human kidney tissue, human liver cancer tissue, mouse eye tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
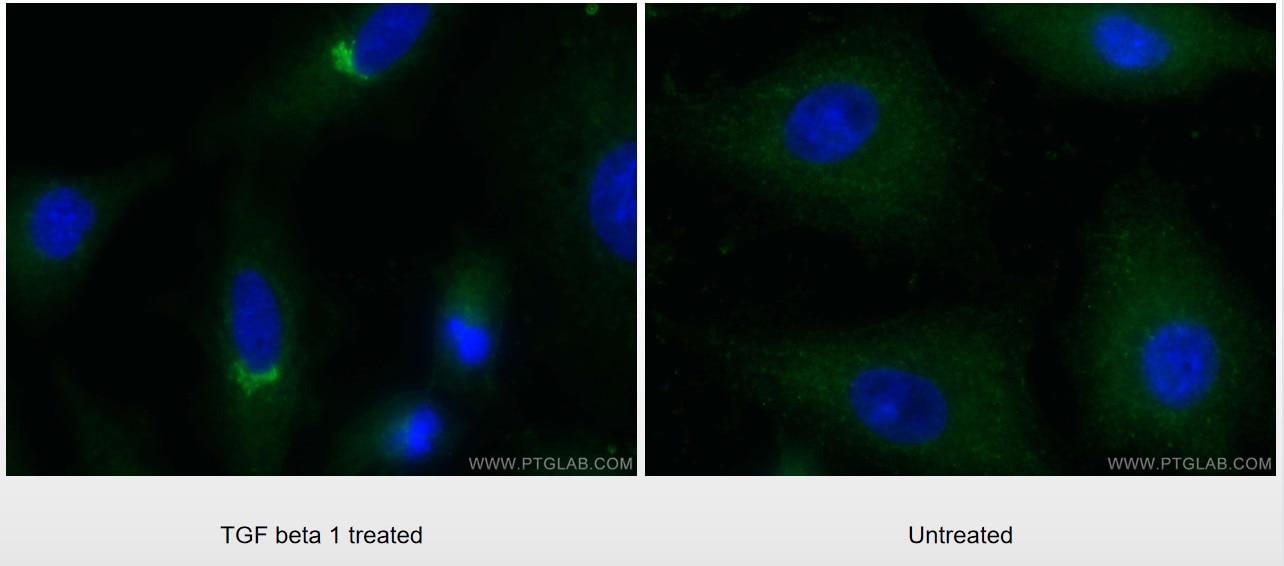
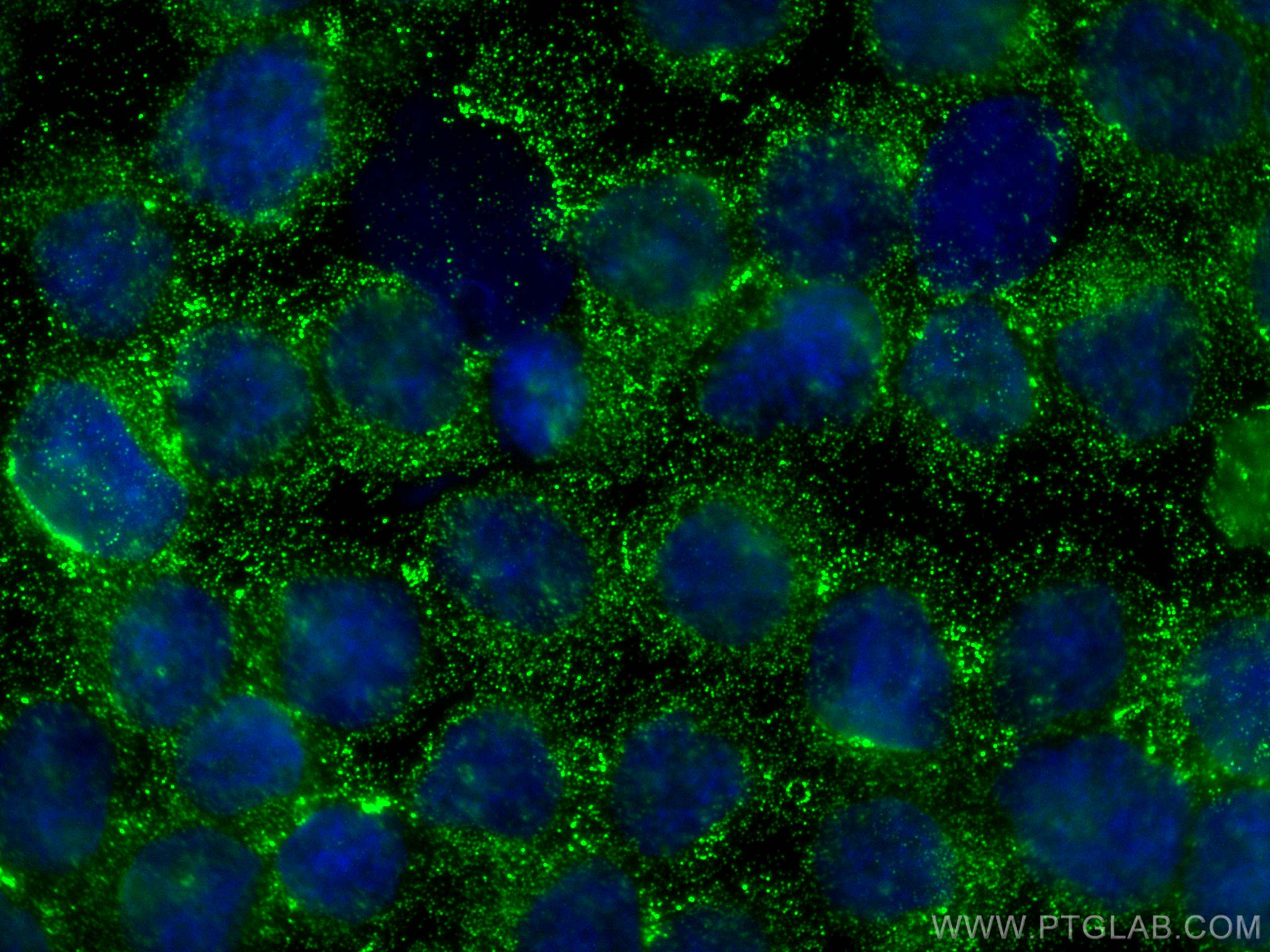
| Positive IF/ICC detected in | TGF beta 1 treated A549 cells, Y79 cells |
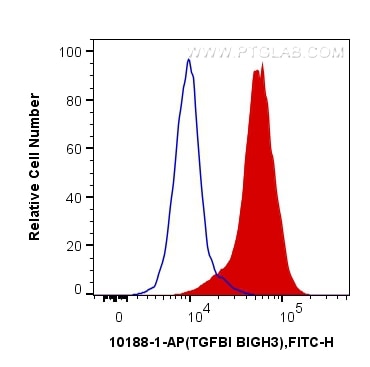
| Positive FC (Intra) detected in | Y79 cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:1000-1:4000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Immunohistochemistry (IHC) | IHC : 1:50-1:500 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:200-1:800 |
| Flow Cytometry (FC) (INTRA) | FC (INTRA) : 0.40 ug per 10^6 cells in a 100 µl suspension |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 9 publications below |
| WB | See 46 publications below |
| IHC | See 37 publications below |
| IF | See 17 publications below |
| IP | See 4 publications below |
Product Information
10188-1-AP targets TGFBI/BIGH3 in WB, IHC, IF/ICC, FC (Intra), IP, Neutralization, ELISA, Cell treatment applications and shows reactivity with human, mouse samples.
| Tested Reactivity | human, mouse |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag0241 Product name: Recombinant human BIGH3 protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 199-406 aa of BC000097 Sequence: NIQIHHYPNGIVTVNCARLLKADHHATNGVVHLIDKVISTITNNIQQIIEIEDTFETLRAAVAASGLNTMLEGNGQYTLLAPTNEAFEKIPSETLNRILGDPEALRDLLNNHILKSAMCAEAIVAGLSVETLEGTTLEVGCSGDMLTINGKAIISNKDILATNGVIHYIDELLIPDSAKTLFELAAESDVSTAIDLFRQAGLGNHLSG Predict reactive species |
| Full Name | transforming growth factor, beta-induced, 68kDa |
| Calculated Molecular Weight | 683 aa, 75 kDa |
| Observed Molecular Weight | 64 kDa |
| GenBank Accession Number | BC000097 |
| Gene Symbol | TGFBI |
| Gene ID (NCBI) | 7045 |
| ENSEMBL Gene ID | ENSG00000120708 |
| RRID | AB_2202311 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity purification |
| UNIPROT ID | Q15582 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
TGFBI, also named as BIGH3, Kerato-epithelin and RGD-CAP, binds to type I, II, and IV collagens. TGFBI is an adhesion protein which may play an important role in cell-collagen interactions. In cartilage, it may be involved in endochondral bone formation. TGFBI is an extracellular matrix adaptor protein, it has been reported to be differentially expressed in transformed tissues. TGFBI is a predictive factor of the response to chemotherapy, and suggest the use of TGFBI-derived peptides as possible therapeutic adjuvants for the enhancement of responses to chemotherapy.(PMID:20509890) Defects in TGFBI are the cause of epithelial basement membrane corneal dystrophy (EBMD). Defects in TGFBI are the cause of corneal dystrophy Groenouw type 1 (CDGG1). Defects in TGFBI are the cause of corneal dystrophy lattice type 1 (CDL1). Defects in TGFBI are a cause of corneal dystrophy Thiel-Behnke type (CDTB). Defects in TGFBI are the cause of Reis-Buecklers corneal dystrophy (CDRB). Defects in TGFBI are the cause of lattice corneal dystrophy type 3A (CDL3A). Defects in TGFBI are the cause of Avellino corneal dystrophy (ACD).
Protocols
| Product Specific Protocols | |
|---|---|
| FC protocol for TGFBI/BIGH3 antibody 10188-1-AP | Download protocol |
| IF protocol for TGFBI/BIGH3 antibody 10188-1-AP | Download protocol |
| IHC protocol for TGFBI/BIGH3 antibody 10188-1-AP | Download protocol |
| IP protocol for TGFBI/BIGH3 antibody 10188-1-AP | Download protocol |
| WB protocol for TGFBI/BIGH3 antibody 10188-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Nat Cell Biol Supermeres are functional extracellular nanoparticles replete with disease biomarkers and therapeutic targets. | ||
Sci Adv Acetylated K676 TGFBIp as a severity diagnostic blood biomarker for SARS-CoV-2 pneumonia. | ||
Genes Dev Extracellular matrix protein betaig-h3/TGFBI promotes metastasis of colon cancer by enhancing cell extravasation.
| ||
Biomaterials Dual peptide-dendrimer conjugate inhibits acetylation of transforming growth factor β-induced protein and improves survival in sepsis. |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
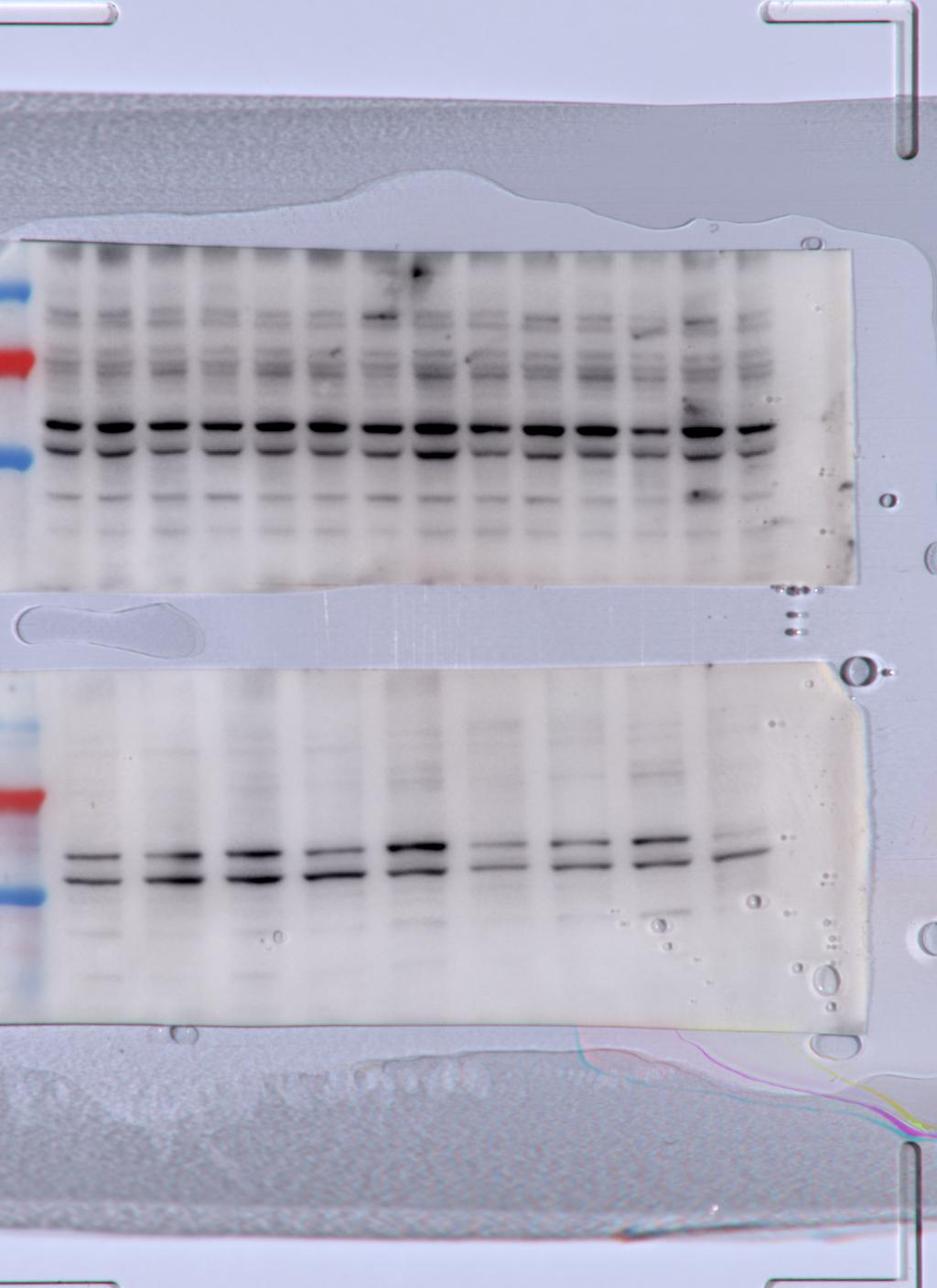
FH Carmelo (Verified Customer) (04-25-2022) | I tested the abs 1:1000 ON at 4C on 40ug spinal cord protein lysates. unfortunately I was not able to distinguish a proper band at the expected size. I will run further optimisation in the incoming weeks.
 |