Tested Applications
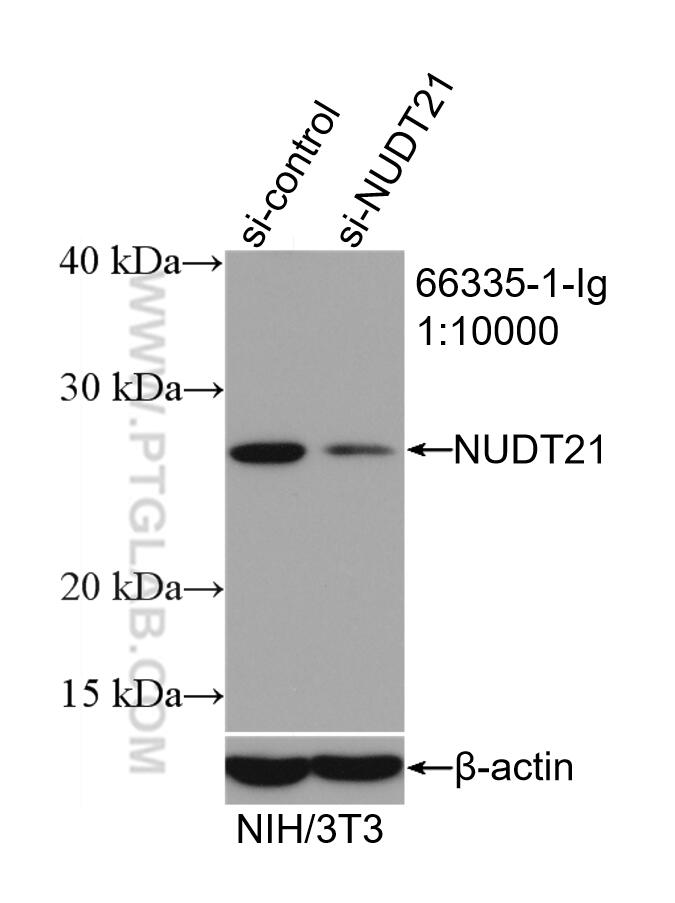
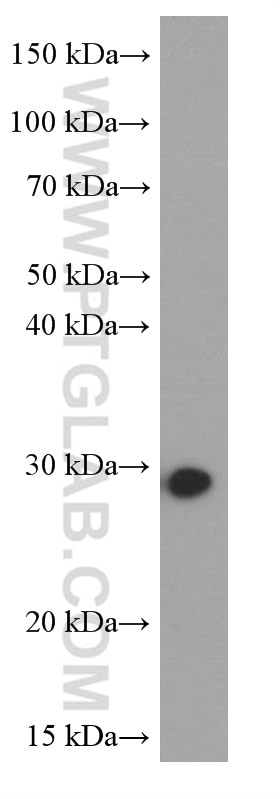
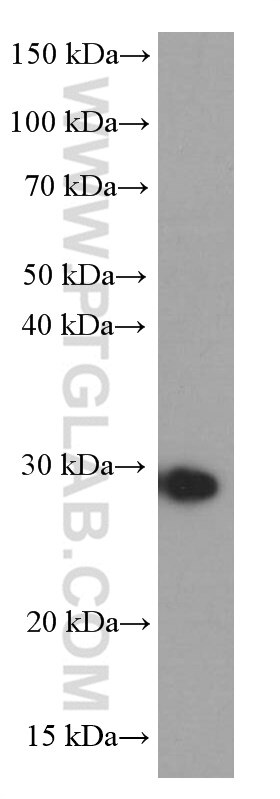
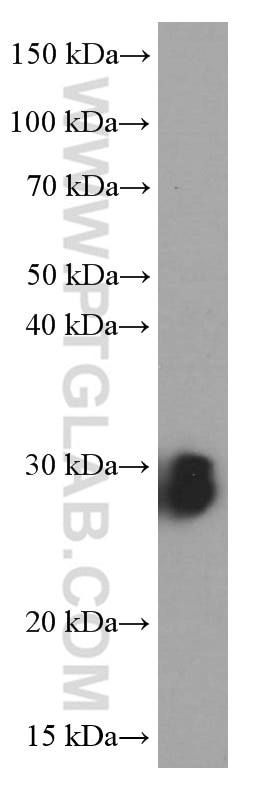
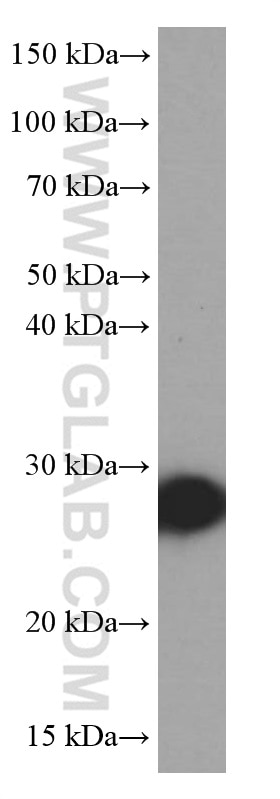
| Positive WB detected in | NIH/3T3 cells, HEK-293 cells, HeLa cells, MCF-7 cells |
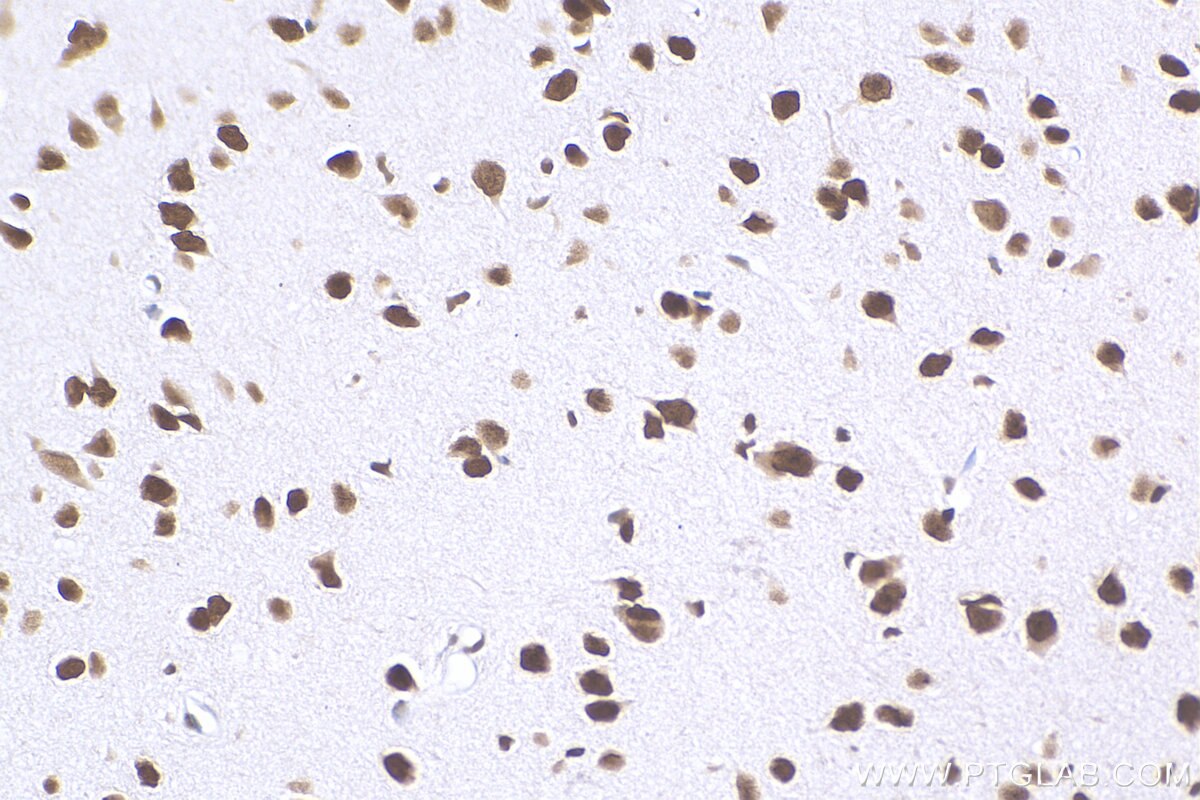
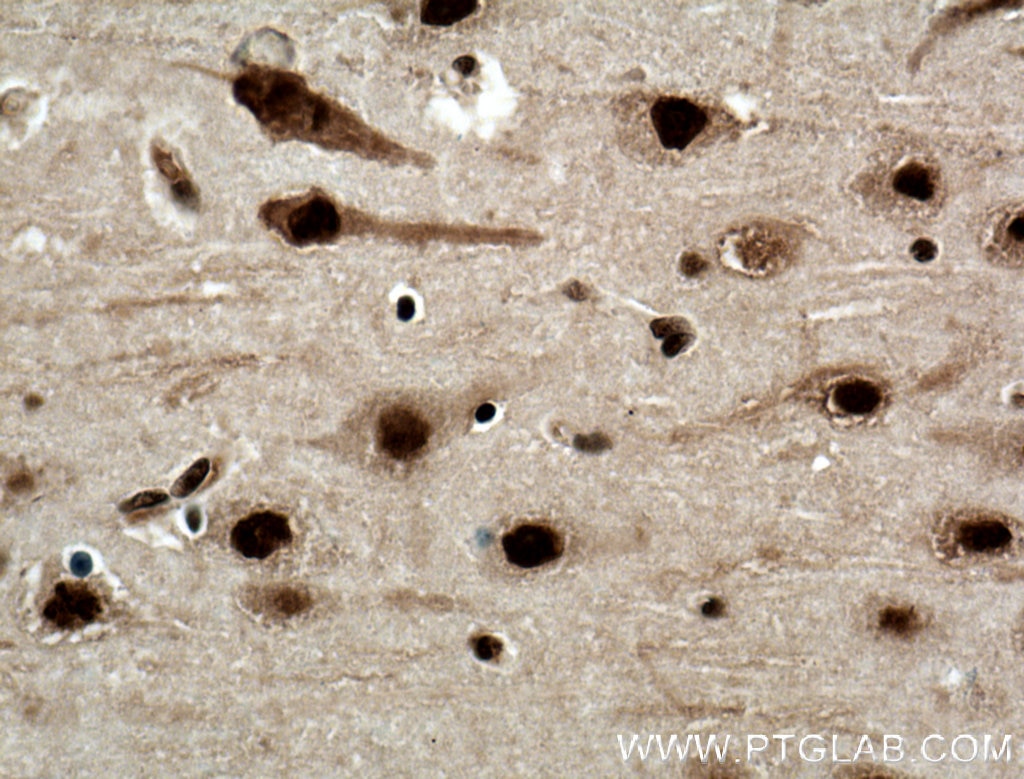
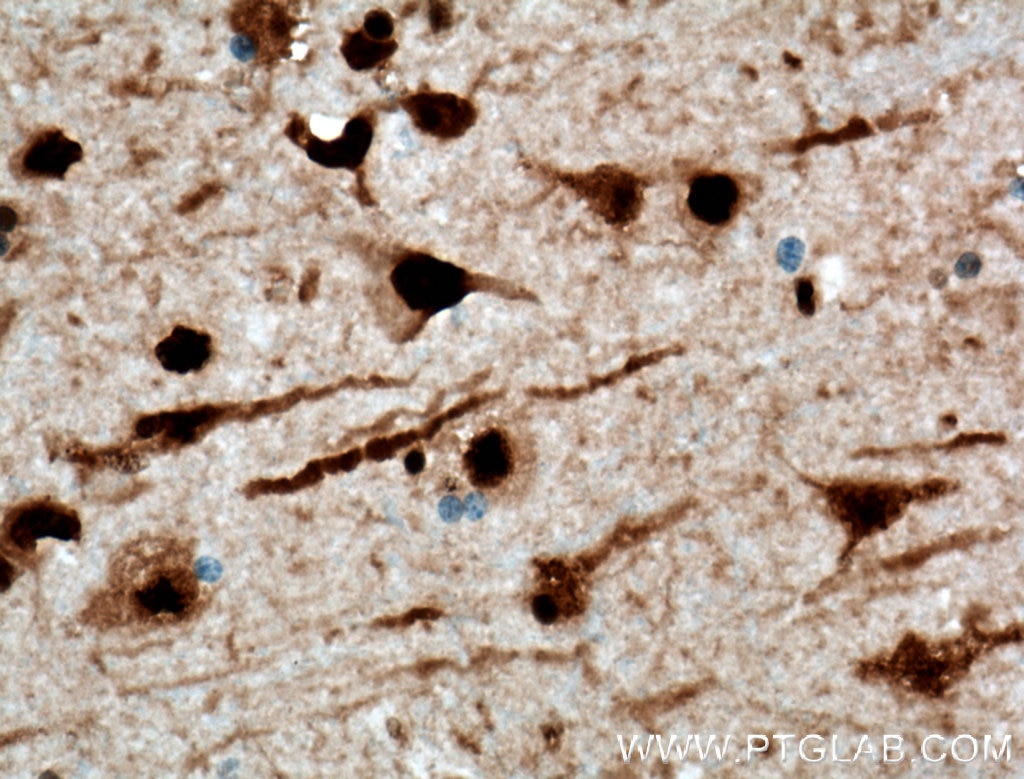
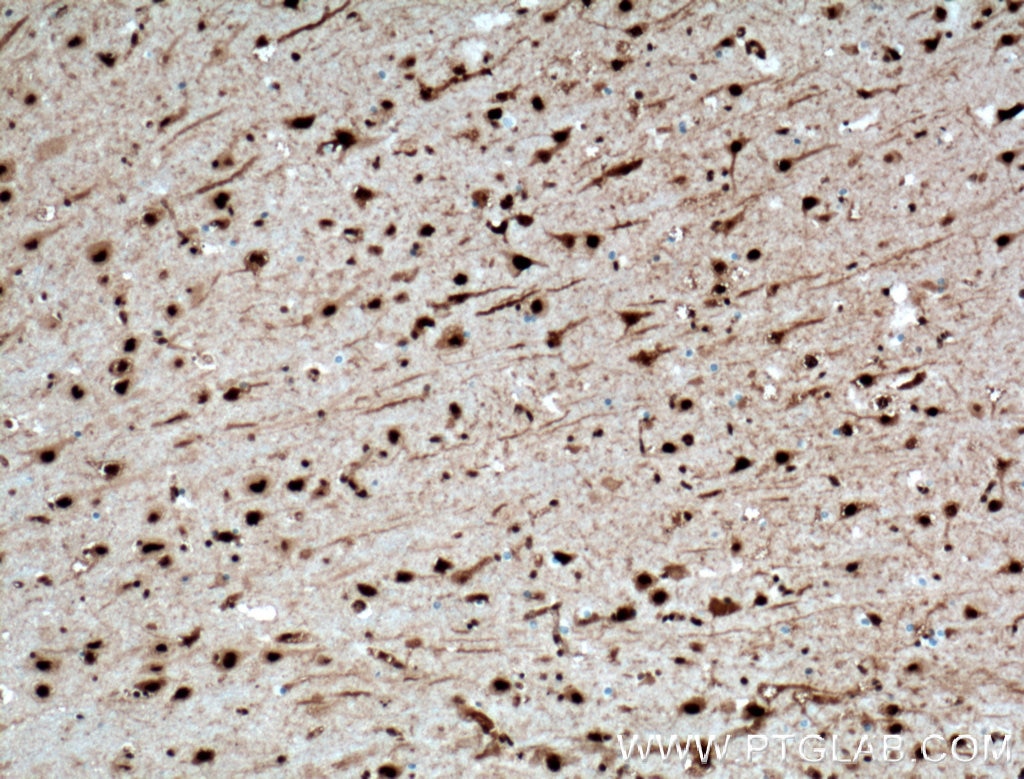
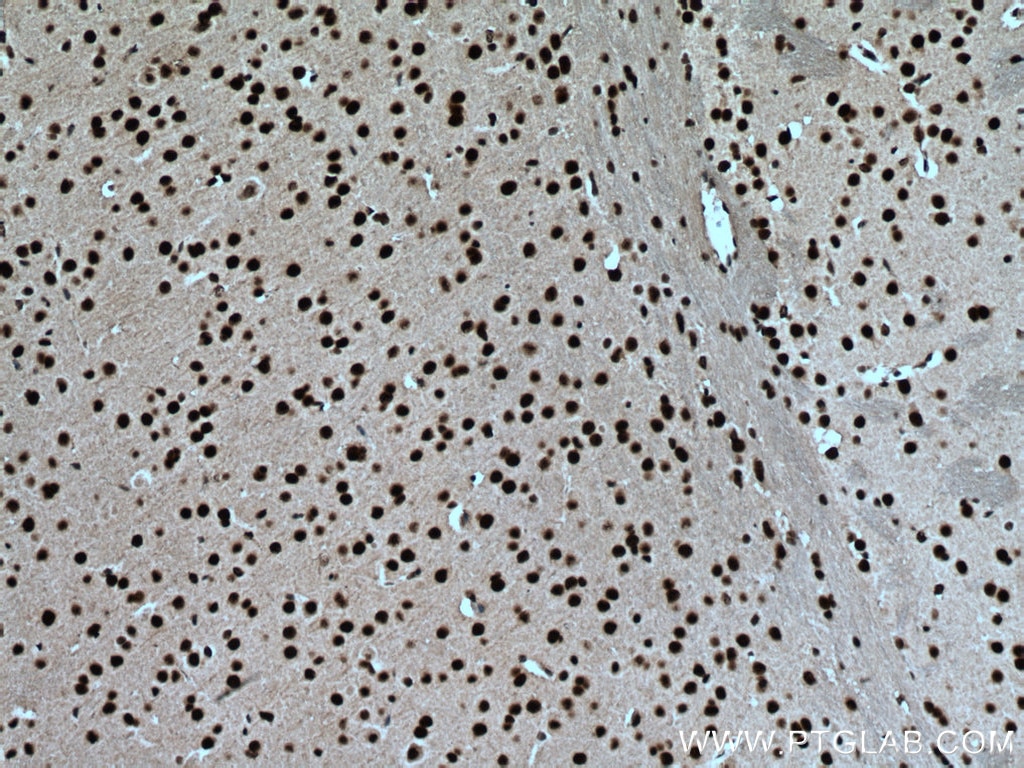
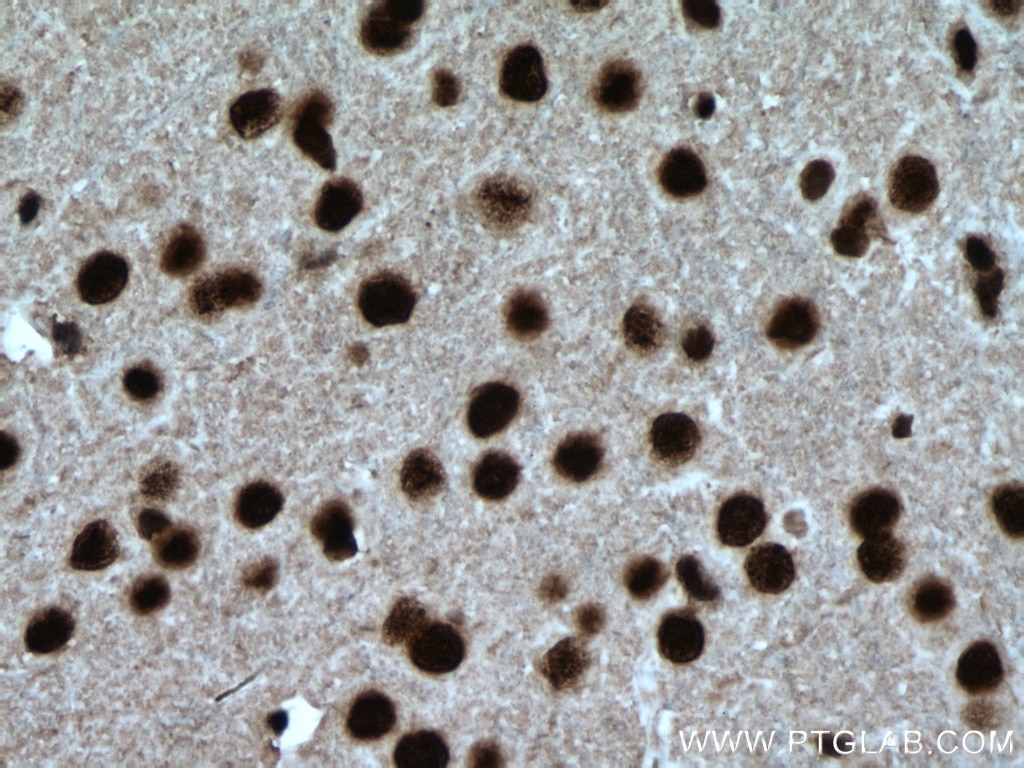
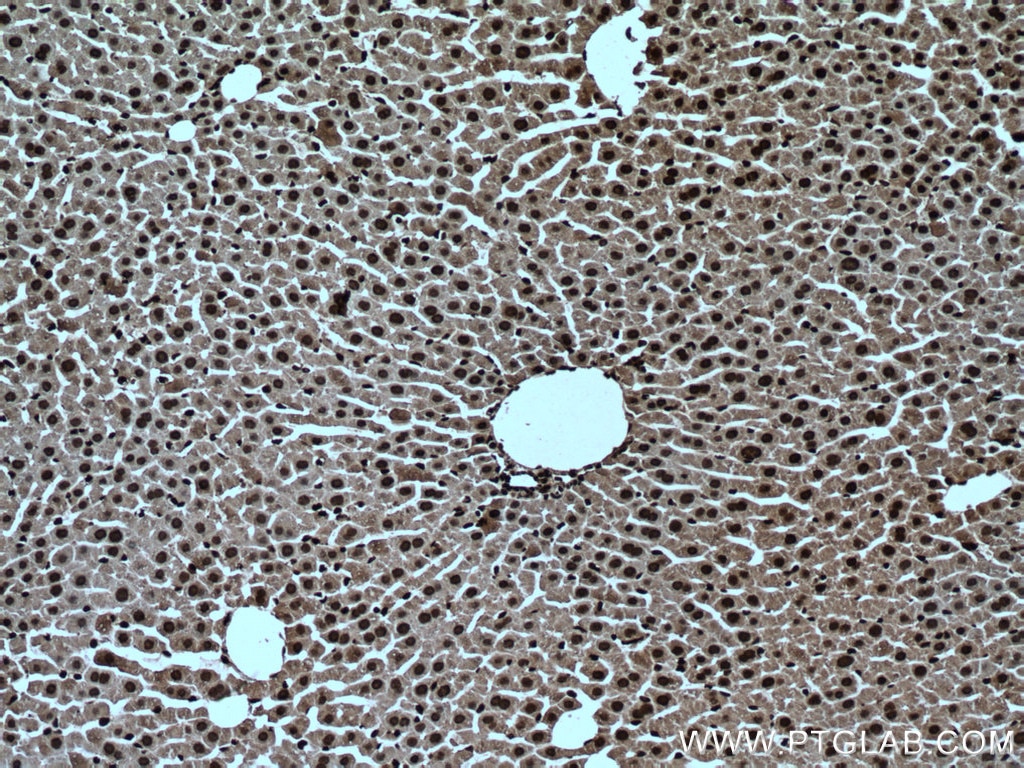
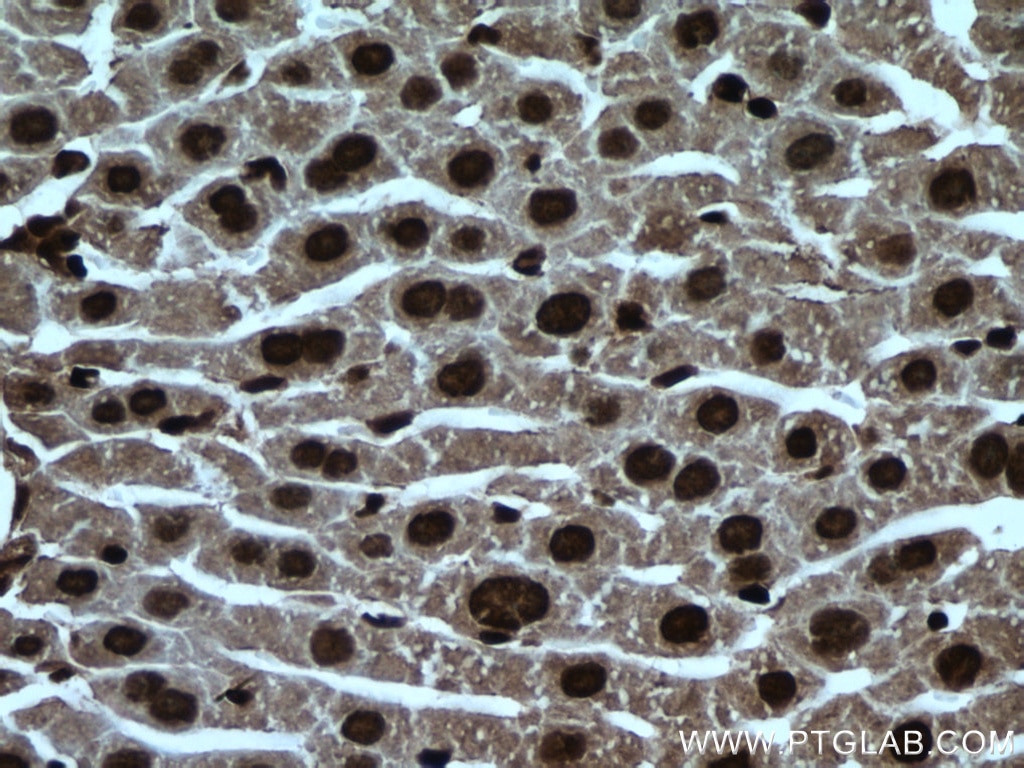
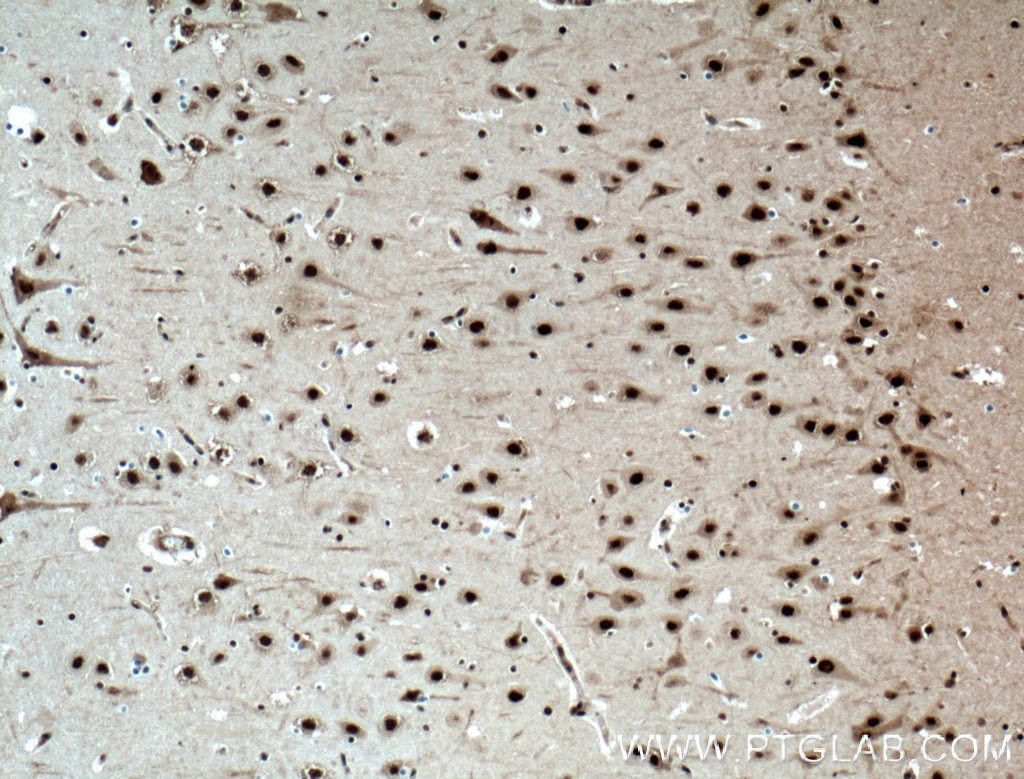
| Positive IHC detected in | human brain tissue, mouse brain tissue, mouse liver tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
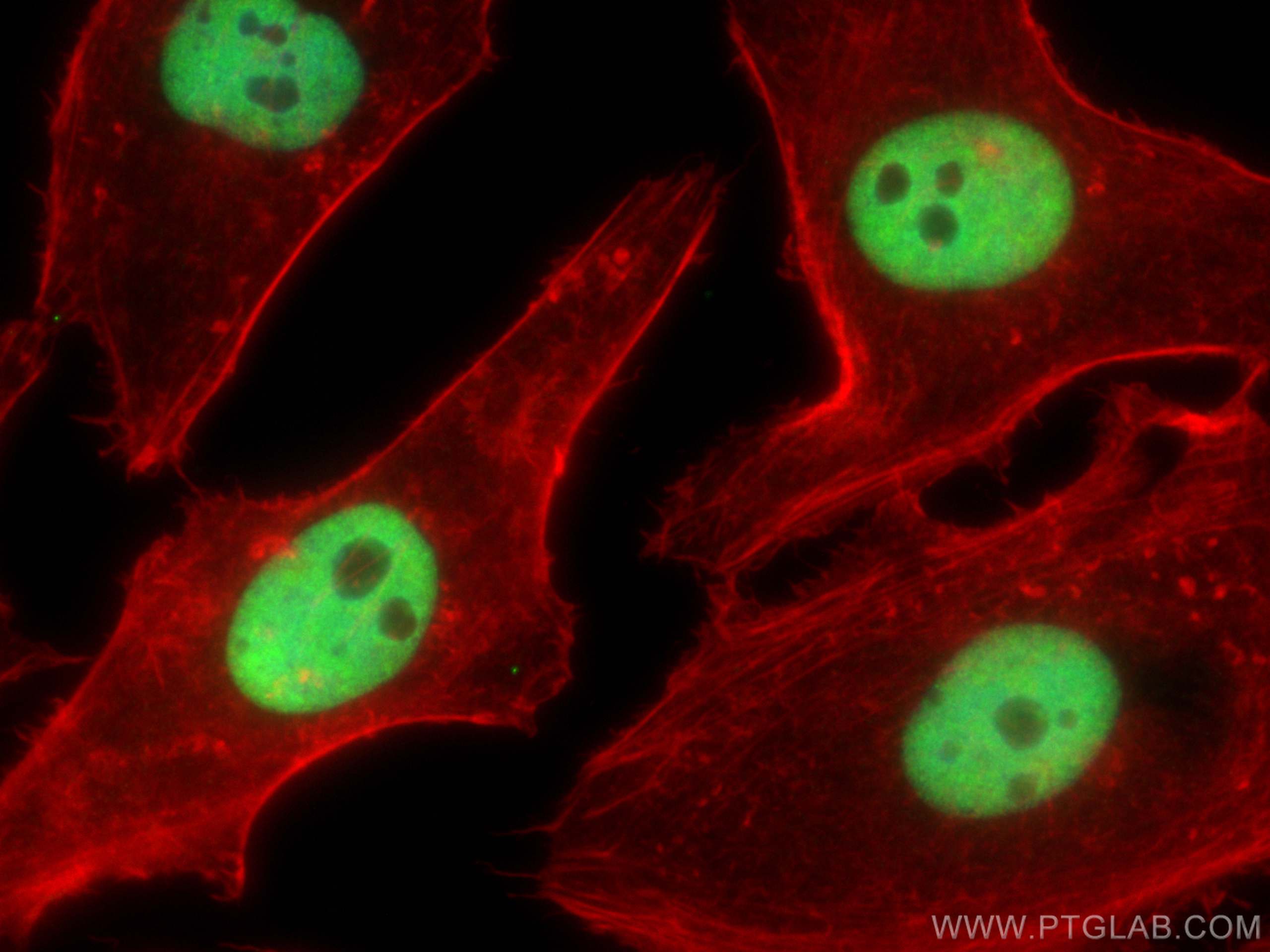
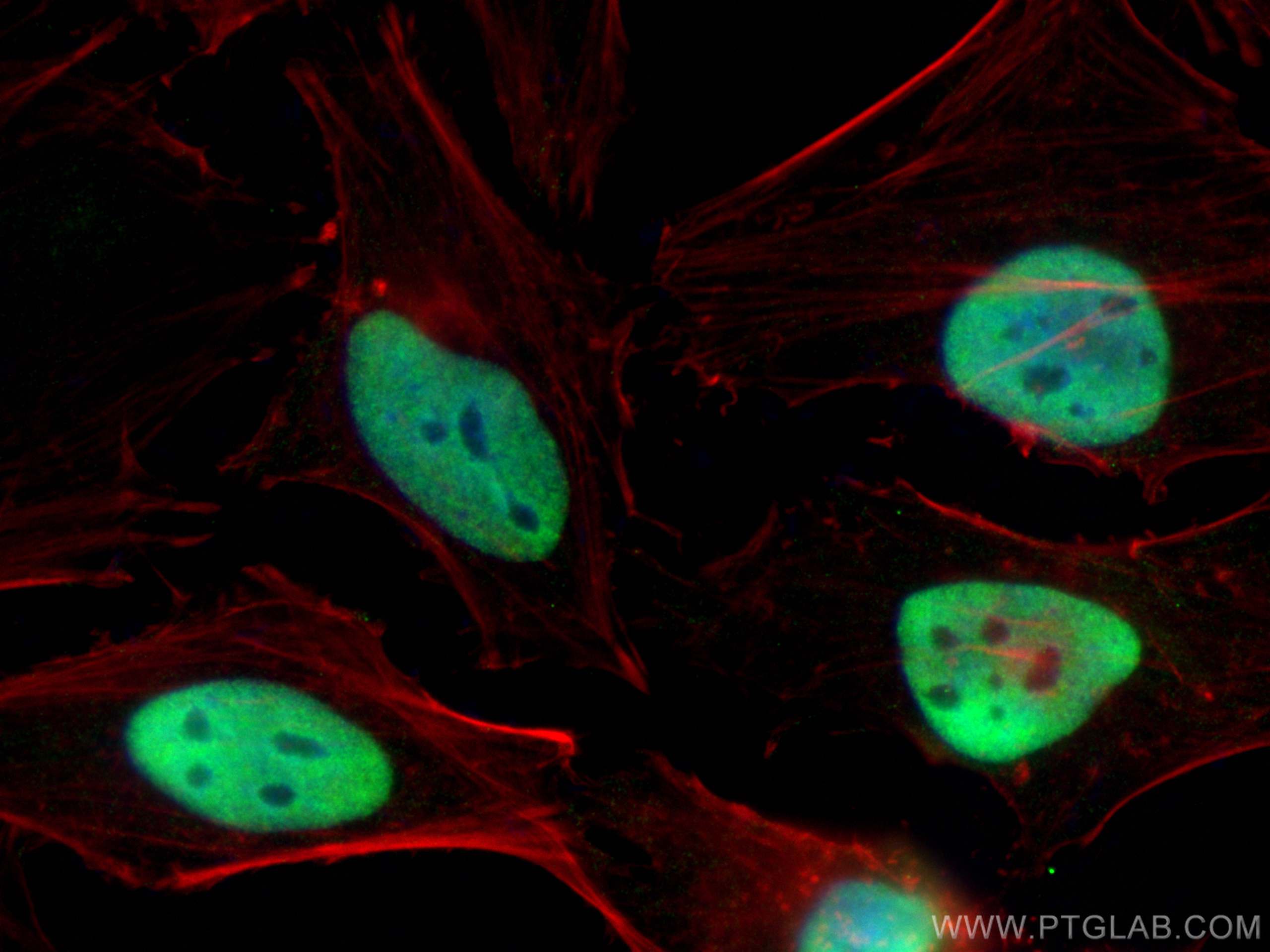
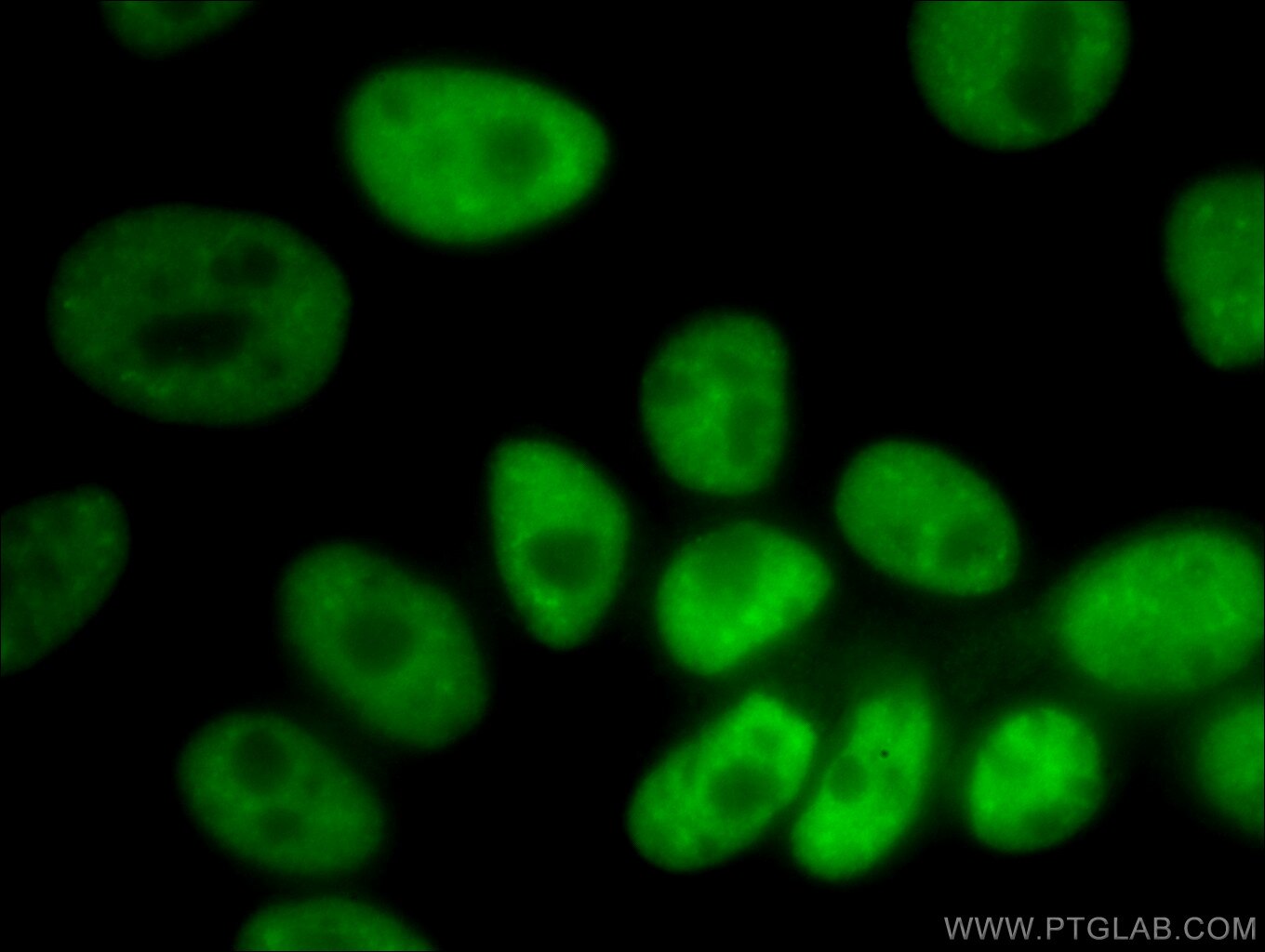
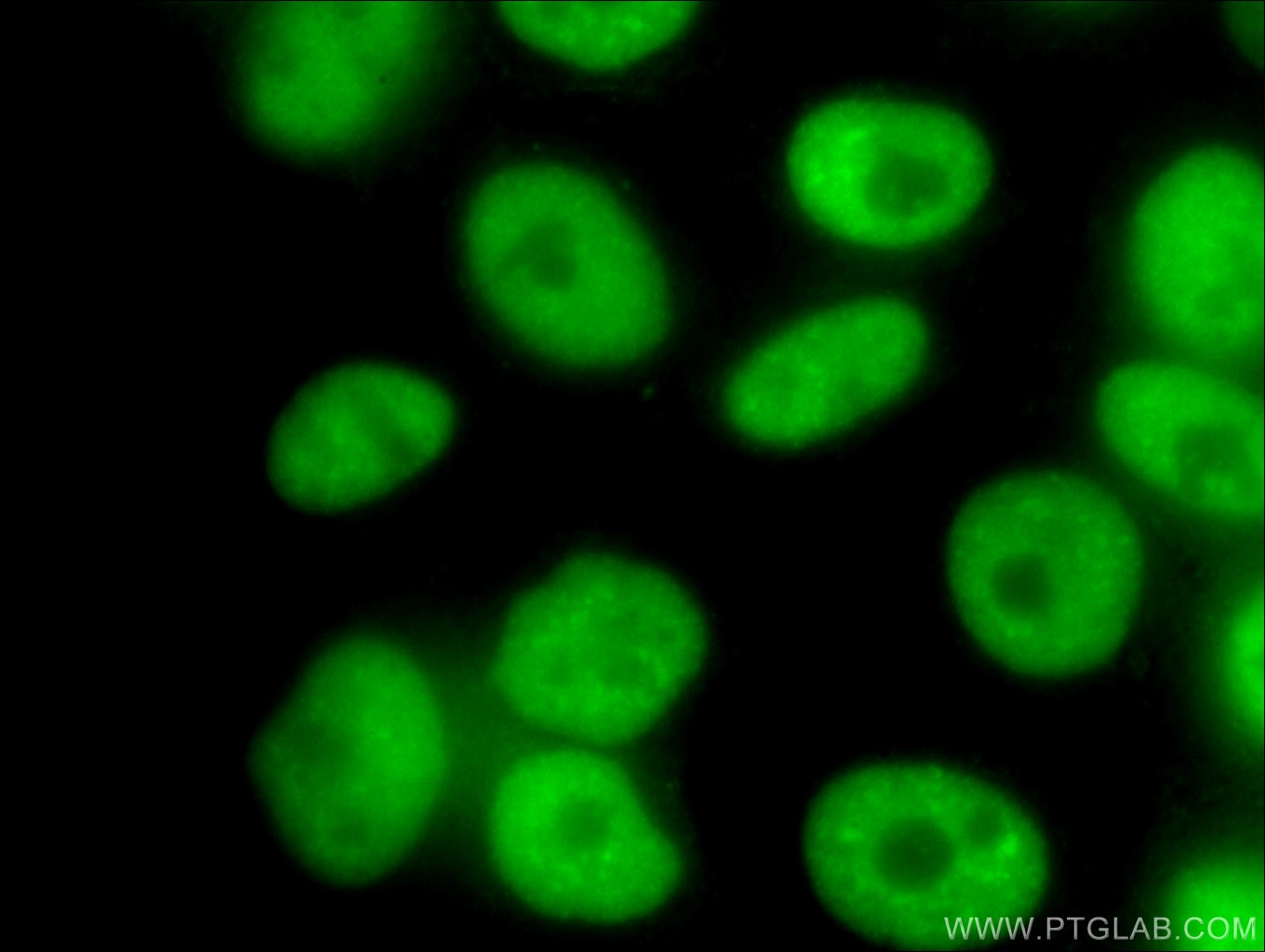
| Positive IF/ICC detected in | HeLa cells, HepG2 cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:1000-1:6000 |
| Immunohistochemistry (IHC) | IHC : 1:500-1:2000 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:1000-1:4000 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| KD/KO | See 1 publications below |
| WB | See 7 publications below |
| IF | See 3 publications below |
| IP | See 2 publications below |
| RIP | See 2 publications below |
Product Information
66335-1-Ig targets NUDT21 in WB, IHC, IF/ICC, IP, RIP, ELISA applications and shows reactivity with human, mouse samples.
| Tested Reactivity | human, mouse |
| Cited Reactivity | human, mouse |
| Host / Isotype | Mouse / IgG1 |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag12260 Product name: Recombinant human NUDT21 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 1-222 aa of BC001403 Sequence: MSVVPPNRSQTGWPRGVTQFGNKYIQQTKPLTLERTINLYPLTNYTFGTKEPLYEKDSSVAARFQRMREEFDKIGMRRTVEGVLIVHEHRLPHVLLLQLGTTFFKLPGGELNPGEDEVEGLKRLMTEILGRQDGVLQDWVIDDCIGNWWRPNFEPPQYPYIPAHITKPKEHKKLFLVQLQEKALFAVPKNYKLVAAPLFELYDNAPGYGPIISSLPQLLSRF Predict reactive species |
| Full Name | nudix (nucleoside diphosphate linked moiety X)-type motif 21 |
| Calculated Molecular Weight | 26 kDa |
| Observed Molecular Weight | 27 kDa |
| GenBank Accession Number | BC001403 |
| Gene Symbol | NUDT21 |
| Gene ID (NCBI) | 11051 |
| RRID | AB_2881715 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein G purification |
| UNIPROT ID | O43809 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
Cleavage and polyadenylation specific factor 5, 25 kDa (CPSF5, synonym: CFIM25) is one subunit of a cleavage factor required for 3' RNA cleavage and polyadenylation processing. The interaction of CPSF5 with the RNA is one of the earliest steps in the assembly of the 3' end processing complex and facilitates the recruitment of other processing factors.
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for NUDT21 antibody 66335-1-Ig | Download protocol |
| IHC protocol for NUDT21 antibody 66335-1-Ig | Download protocol |
| WB protocol for NUDT21 antibody 66335-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Mol Cell A Cancer-Specific Ubiquitin Ligase Drives mRNA Alternative Polyadenylation by Ubiquitinating the mRNA 3' End Processing Complex. | ||
Adv Sci (Weinh) NUDT21-mediated Alternative Polyadenylation of CDK19 Reprograms Cholesterol Biosynthesis to Drive Colorectal Cancer Progression.
| ||
Cell Rep LINC00921 reduces lung cancer radiosensitivity by destabilizing NUDT21 and driving aberrant MED23 alternative polyadenylation | ||
J Mol Cell Biol U1 snRNP proteins promote proximal alternative polyadenylation sites by directly interacting with 3' end processing core factors | ||
Viruses HIV-1 Capsid Rapidly Induces Long-Lived CPSF6 Puncta in Non-Dividing Cells, but Similar Puncta Already Exist in Uninfected T-Cells | ||
Cell Rep CPSF1 inhibition promotes widespread use of intergenic polyadenylation sites and impairs glycolysis in prostate cancer cells |