GFP (green fluorescent protein): Properties, origin, specifications, tips
Everything you need to know about GFP: properties and applications
Summary
- GFP (green fluorescent protein) is a fluorescent protein originally discovered in the jellyfish Aequorea victoria and is widely used as a live-cell reporter and protein tag in research.
- GFP works as a fusion tag — by genetically attaching GFP to a protein of interest, researchers can track its location, movement, and interactions in real time using fluorescence microscopy.
- EGFP (enhanced GFP) is the most commonly used derivative, with an excitation peak at 488 nm and emission at 507 nm, and a molecular weight of 26.9 kDa.
- GFP and its derivatives are widely used across applications including fluorescence microscopy, reporter assays, immunoprecipitation, protein purification, and FRET-based signal transduction studies.
- Anti-GFP Nanobodies like the GFP-Trap offer a highly specific and low-background method for pulling down GFP-tagged proteins in immunoprecipitation experiments.
What is GFP?
GFP stands for green fluorescent protein. GFP is a fluorescent protein that can be expressed in vivo. If GFP is exposed to light, it emits a green fluorescent signal. This property has had an enormous impact on cell biology by enabling the imaging of almost any protein, in transcription studies by working as a reporter gene, and in biochemical applications.
Origin of GFP
GFP is an endogenous protein from the jellyfish Aequorea Victoria. It was isolated by Osamu Shimomura in 1962. In 1992, the sequence of GFP was cloned (Douglas Prasher) and Martin Chalfie’s lab expressed the sequence in vivo. Roger Tsien’s lab improved GFP and managed to convert it into a commonly used research tool. In 2008, The Nobel Prize in Chemistry was awarded “for the discovery and development of the green fluorescent protein, GFP.” See Roger Tsien’s Nobel Prize lecture here.
How does GFP work?
GFP is a protein and, like other proteins, it can be expressed by living organisms. Once GFP is expressed and properly folded, it shows fluorescent properties. If excited by light in the ultra-violet or blue spectrum, GFP emits green light (for more details please see sections 8 and 9). This property has had an enormous impact on cell biology: GFP and proteins fused to GFP can be detected as GFP works as a fluorescent tag. This has actually made lots of new experiments possible.
What are GFP-tagged proteins and GFP-fusion proteins
Endogenous proteins do not contain protein or peptide tags and therefore are sometimes difficult to detect in an assay. One solution that enables easy detection is to genetically fuse protein and peptide tags to the protein of interest. Tagged proteins can be used for purposes such as immunoprecipitation, microscopy, protein purification, Western blotting, protein arrays, etc. Proteins fused to GFP are called GFP-tagged proteins or GFP-fusion proteins. GFP-tagged proteins are often used for fluorescence microscopy, immunoprecipitation, protein purification, and Western blotting.
GFP emission spectrum and excitation peaks
| Excitation max | Emission max | |
|---|---|---|
| GFP | 395 nm and 475 nm | 509 nm |
| EGFP (most common derivative) | 488 nm | 507 nm |
For more details see https://www.fpbase.org/protein/avgfp/ and https://www.fpbase.org/protein/egfp/
Which applications can GFP be used for?
GFP has been used in many different applications such as:
- imaging of proteins (epi-fluorescent microscopy, confocal microscopy, super-resolution microscopy)
- reporter assays (transcription studies)
- signal transduction studies (FRET: fluorescence resonance energy transfer)
- biochemical applications (immunoprecipitation, protein purification)
- a bio-sensor (pH, calcium)
GFP derivatives
Many fluorescent proteins are based on the GFP sequence. These fluorescent proteins are genetically engineered and have different properties such as different excitation/emission spectra, photo or pH-stability, folding properties, half-time etc. Here is a rough overview:
| BFP | GFP Envy | Superfolder GFP (sfGFP) |
| CFP | GFP S65T | yeast EGFP |
| AcGFP | GFPSpark | Citrine |
| Clover | GFPuv | Ecitrine |
| EGFP | mClover (Clover A260K) | EYFP |
| Emerald | mEGFP | Venus |
| G3GFP | mGFP (monomeric GFP) | YFP |
| GFP | Monomeric EGFP A206K | Ypet |
| GFP (cycle 3) | mPhluorin | |
| GFP5 | PA-GFP |
See https://www.fpbase.org/protein/avgfp/ for the complete list.
Structure of GFP
GFP and GFP derivate EGFP have a β-barrel structure. In the center of this β-barrel, there are 3 amino acids. These acids and the cyclization and oxidation of their backbone form a two-ring chromophore.
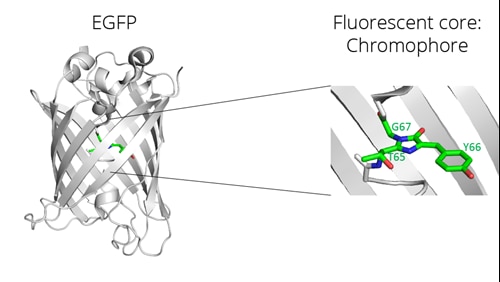
Left: beta-barrel structure and fluorescent core of enhanced GFP (EGFP), a GFP derivate; right: magnification of the fluorescent core, the two-ring chromophore
Why is GFP fluorescent?
The two-ring chromophore of GFP absorbs and emits light, e.g., light photons, in the visible green spectrum. The chromophore of GFP lies in the center of a beta-barrel structure. The two-ring chromophore is formed by oxidation and cyclization of the backbone of 3 amino acids: Threonine 65, Tyrosine 66, and Glycine 67. This process occurs during the folding of the protein and depends on different factors such as pH, temperature, and oxygen concentration.
Properties and Size of EGFP
Size:
Number of amino acids: 239
Molecular weight (MW): 26,9 kDa
Properties:
Extinction coefficient (EC): 55,900 M-1 cm-1
Maturation rate (at 37°C): 25 min
EGFP Sequences
EGFP amino acid sequence:
| 10 | 20 | 30 | 40 | 50 |
| MVSKGEELFT | GVVPILVELD | GDVNGHKFSV | SGEGEGDATY | GKLTLKFICT |
| 60 | 70 | 80 | 90 | 100 |
| TGKLPVPWPT | LVTTLTYGVQ | CFSRYPDHMK | QHDFFKSAMP | EGYVQERTIF |
| 110 | 120 | 130 | 140 | 150 |
| FKDDGNYKTR | AEVKFEGDTL | VNRIELKGID | FKEDGNILGH | KLEYNYNSHN |
| 160 | 170 | 180 | 190 | 200 |
| VYIMADKQKN | GIKVNFKIRH | NIEDGSVQLA | DHYQQNTPIG | DGPVLLPDNH |
| 210 | 220 | 230 | ||
| YLSTQSALSK | DPNEKRDHMV | LLEFVTAAGI | TLGMDELYK |
Source: https://www.uniprot.org/uniprot/C5MKY7
EGFP DNA sequences:
| atggtgagca | agggcgagga | gctgttcacc | ggggtggtgc | ccatcctggt | cgagctggac | 60 |
| ggcgacgtaa | acggccacaa | gttcagcgtg | tccggcgagg | gcgagggcga | tgccacctac | 120 |
| ggcaagctga | ccctgaagtt | catctgcacc | accggcaagc | tgcccgtgcc | ctggcccacc | 180 |
| ctcgtgacca | ccctgaccta | cggcgtgcag | tgcttcagcc | gctaccccga | ccacatgaag | 240 |
| cagcacgact | tcttcaagtc | cgccatgccc | gaaggctacg | tccaggagcg | caccatcttc | 300 |
| ttcaaggacg | acggcaacta | caagacccgc | gccgaggtga | agttcgaggg | cgacaccctg | 360 |
| gtgaaccgca | tcgagctgaa | gggcatcgac | ttcaaggagg | acggcaacat | cctggggcac | 420 |
| aagctggagt | acaactacaa | cagccacaac | gtctatatca | tggccgacaa | gcagaagaac | 480 |
| ggcatcaagg | tgaacttcaa | gatccgccac | aacatcgagg | acggcagcgt | gcagctcgcc | 540 |
| gaccactacc | agcagaacac | ccccatcggc | gacggccccg | tgctgctgcc | cgacaaccac | 600 |
| tacctgagca | cccagtccgc | cctgagcaaa | gaccccaacg | agaagcgcga | tcacatggtc | 660 |
| ctgctggagt | tcgtgaccgc | cgccgggatc | actctcggca | tggacgagct | gtacaagtaa | 720 |
Source: https://www.ebi.ac.uk/ena/browser/view/ACS32473
Anti-GFP antibodies and Nanobodies
There are different polyclonal and monoclonal antibodies and an anti-GFP Nanobody available commercially:
- Immunoprecipitation: ChromoTek’s GFP Nanobody GFP-Trap
- Very low background
- No dissociating heavy and light antibody chains
- Stringent washing
- Immunofluorescence: GFP-Booster
- Better tissue penetration
- Higher resolution
- Western blotting: GFP antibody [3H9], rat monoclonal
- Immunofluorescence, Western blotting: GFP antibody [PABG1], rabbit polyclonal
GFP beads
Chromotek offers GFP Nanobodies conjugated to beads for immunoprecipitation and unconjugated GFP Nanobodies/VHHs:
GFP-Trap Agarose: anti-GFP Nanobody conjugated to agarose beads
GFP-Trap Magnetic Agarose: anti-GFP Nanobody conjugated to magnetic agarose beads
GFP-Trap Magnetic Particles M-270: anti-GFP Nanobody conjugated to magnetic particles M-270 for analysis of very large proteins/complexes.
GFP VHH: anti-GFP Nanobody
Free sample
You can test the GFP-Trap yourself. Just request a free sample here:
FAQs
Can GFP antibodies detect YFP, CFP, or other derivatives?
That depends on the antibody. Most GFP antibodies also recognize other derivatives, especially if there are just a few amino acid substitutions compared to GFP or EGFP. For example, the GFP-Trap recognizes AcGFP, Clover, eGFP, Emerald, GFP, GFP5, GFP Envy, GFP, S65T, mGFP, mPhluorin, PA-GFP, Superfolder GFP, TagGFP, TagGFP2, monomeric eGFP K206A, CFP, YFP, Citrine, eCitrine, eYFP, Venus, Ypet, BFP.
Is GFP still fluorescent after fixation?
There is no general answer to this question as it depends on the fixation method and procedure.
Related Content
- What GFP-Trap should I use for my immunoprecipitation?
- GFP-Trap for immunoprecipitation
- How to elute GFP-fusion protein from the GFP-Trap
- How to conduct a Co-immunoprecipitation (Co-IP)
- Mass spec-compatible immunoprecipiation for GFP, mNeonGreen, Myc, RFP, Spot, and TurboGFP
- The best anti-GFP antibody for immunoprecipitation: GFP-Trap
- Green fluorescent protein (GFP) in plant research
- Virus research using GFP
- New limit in protein complex stability: GFP-binding protein:GFP complex
- Antibody Specificity and affinity in Immunoprecipitation
- 5 Tips for better immunoprecipitation
Support
Request a Free Nano-Trap Sample
Nano-Trap system provides fast, reliable, and effective immunoprecipitation of fusion proteins. More than 3,000 peer-reviewed articles have already been published using ChromoTek's Nano-Traps.

