Tested Applications
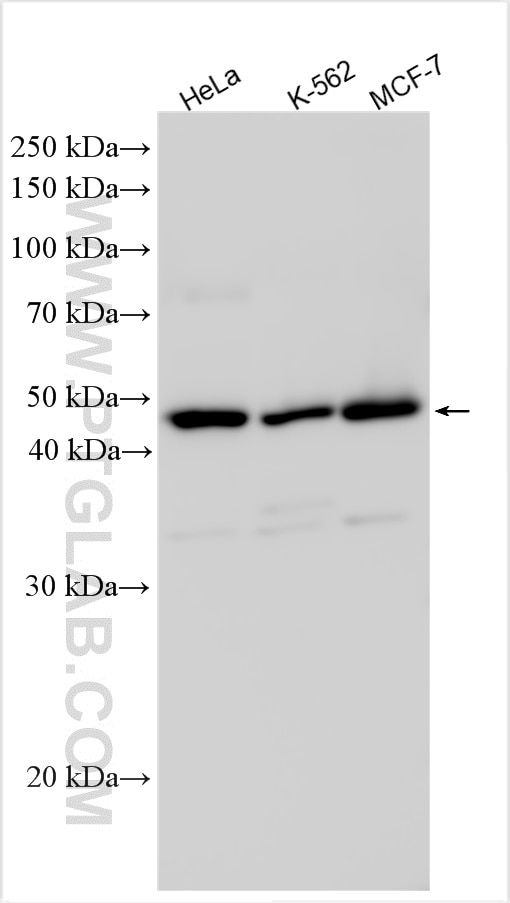
| Positive WB detected in | HeLa cells, K-562 cells, MCF-7 cells |
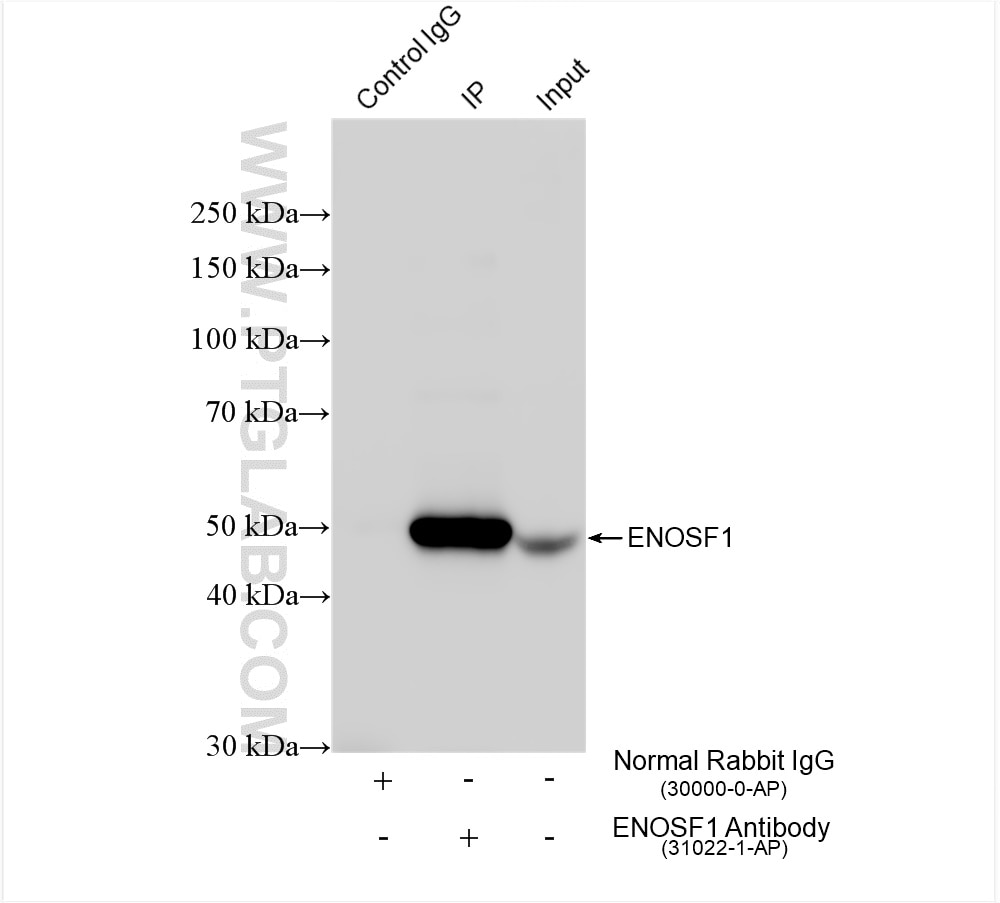
| Positive IP detected in | MCF-7 cells |
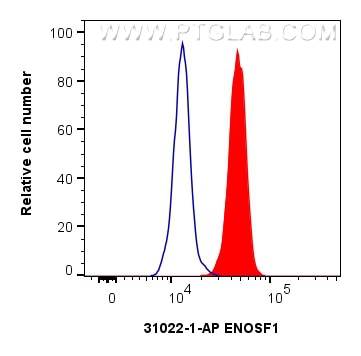
| Positive FC (Intra) detected in | HeLa cells |
Recommended dilution
| Application | Dilution |
|---|---|
| Western Blot (WB) | WB : 1:1000-1:4000 |
| Immunoprecipitation (IP) | IP : 0.5-4.0 ug for 1.0-3.0 mg of total protein lysate |
| Flow Cytometry (FC) (INTRA) | FC (INTRA) : 0.25 ug per 10^6 cells in a 100 µl suspension |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Product Information
31022-1-AP targets ENOSF1 in WB, FC (Intra), IP, ELISA applications and shows reactivity with human samples.
| Tested Reactivity | human |
| Host / Isotype | Rabbit / IgG |
| Class | Polyclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag34665 Product name: Recombinant human ENOSF1 protein Source: e coli.-derived, PGEX-4T Tag: GST Domain: 1-443 aa of BC001285 Sequence: MVRGRISRLSVRDVRFPTSLGGHGADAMHTDPDYSAAYVVIETDAEDGIKGCGITFTLGKGTEVVVCAVNALAHHVLNKDLKDIVGDFRGFYRQLTSDGQLRWIGPEKGVVHLATAAVLNAVWDLWAKQEGKPVWKLLVDMDPRMLVSCIDFRYITDVLTEEDALEILQKGQIGKKEREKQMLAQGYPAYTTSCAWLGYSDDTLKQLCAQALKDGWTRFKVKVGADLQDDMRRCQIIRDMIGPEKTLMMDANQRWDVPEAVEWMSKLAKFKPLWIEEPTSPDDILGHATISKALVPLGIGIATGEQCHNRVIFKQLLQAKALQFLQIDSCRLGSVNENLSVLLMAKKFEIPVCPHAGGVGLCELVQHLIIFDYISVSASLENRVCEYVDHLHEHFKYPVMIQRASYMPPKDPGYSTEMKEESVKKHQYPDGEVWKKLLPAQEN Predict reactive species |
| Full Name | enolase superfamily member 1 |
| Calculated Molecular Weight | 50 kDa |
| Observed Molecular Weight | 49 kDa |
| GenBank Accession Number | BC001285 |
| Gene Symbol | ENOSF1 |
| Gene ID (NCBI) | 55556 |
| RRID | AB_3669818 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Antigen affinity Purification |
| UNIPROT ID | Q7L5Y1 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
ENOSF1 (Mitochondrial enolase superfamily member 1) is also named as RTS and TYMSAS. The ENOSF1 gene, responsible for encoding the protein mitochondrial enolase superfamily member 1 (ENOF1), exhibits three distinct isoforms. One of these isoforms has been identified as an l-fuconate dehydratase, actively participating in the catabolism of L-fucose, a sugar integral to the carbohydrate composition of cellular glycoproteins (PMID: 38203276). The ENOSF1 transcript has been shown to act as antisense RNA and inhibit TYMS expression (PMID: 35931051).
Protocols
| Product Specific Protocols | |
|---|---|
| FC protocol for ENOSF1 antibody 31022-1-AP | Download protocol |
| IP protocol for ENOSF1 antibody 31022-1-AP | Download protocol |
| WB protocol for ENOSF1 antibody 31022-1-AP | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |