Tested Applications
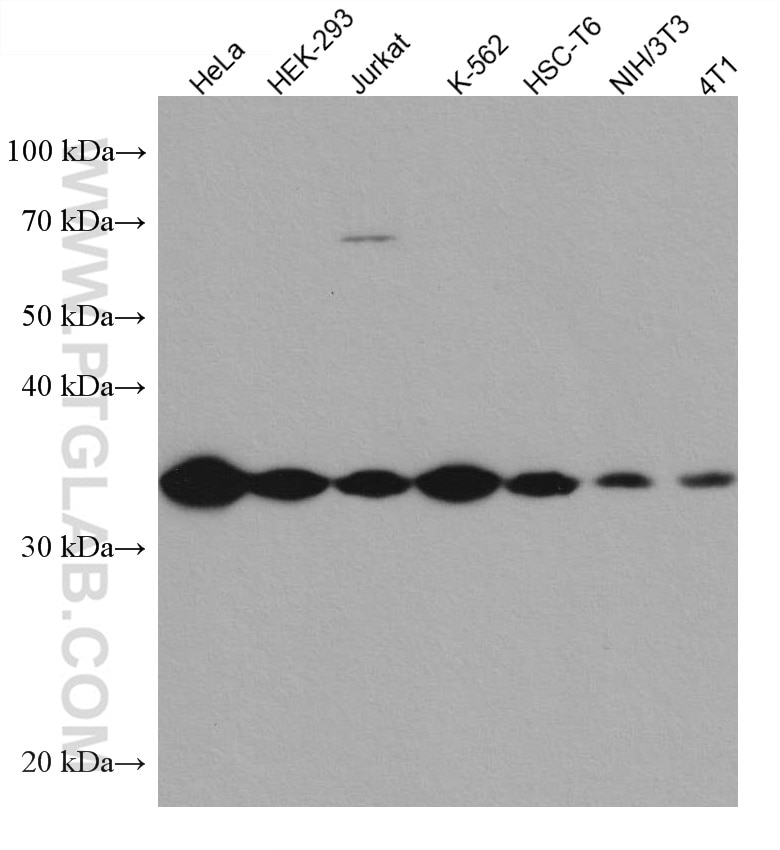
| Positive WB detected in | HeLa cells, HEK-293 cellls, Jurkat cells, K-562 cells, HSC-T6 cells, NIH/3T3 cells, 4T1 cells |
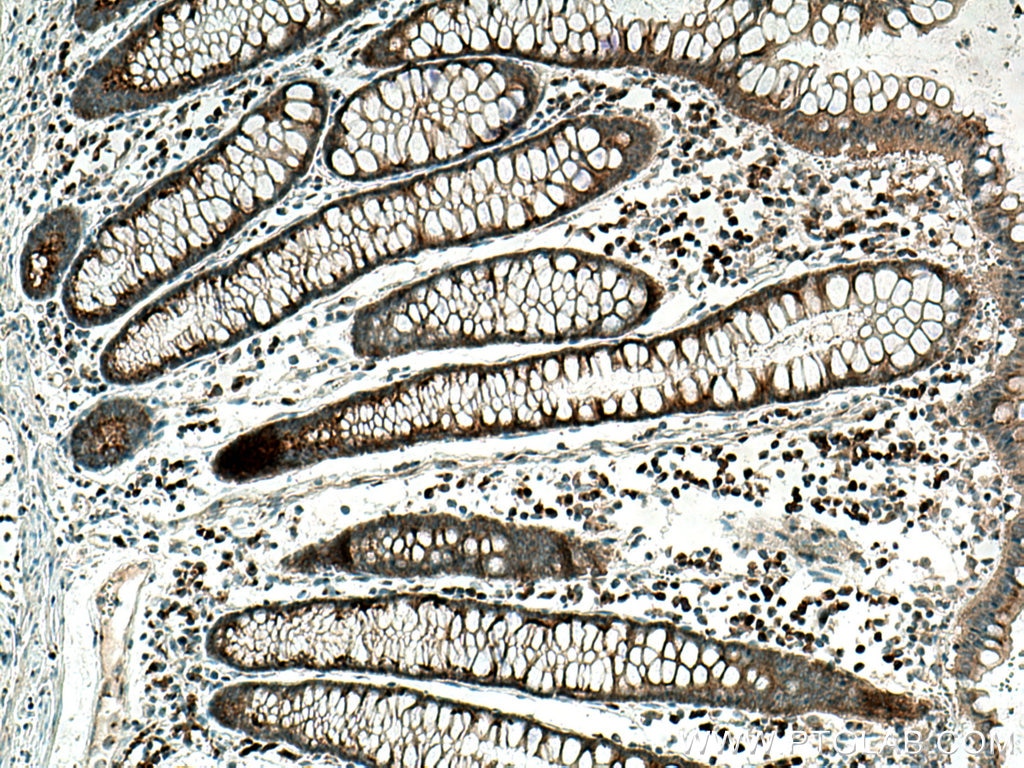
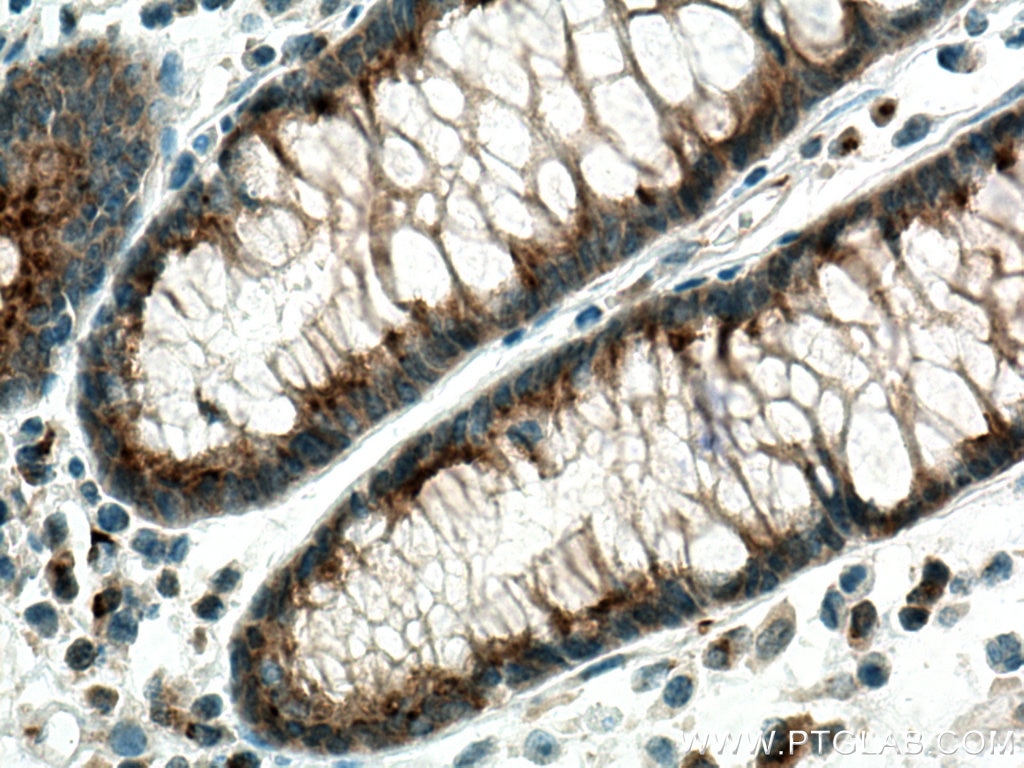
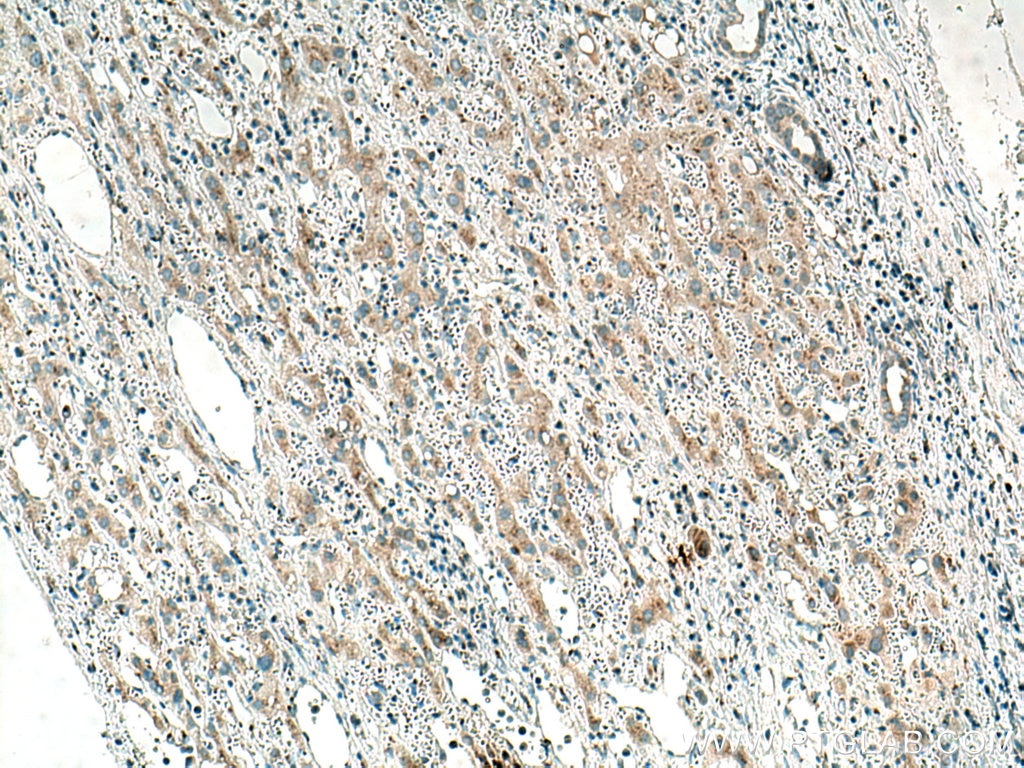
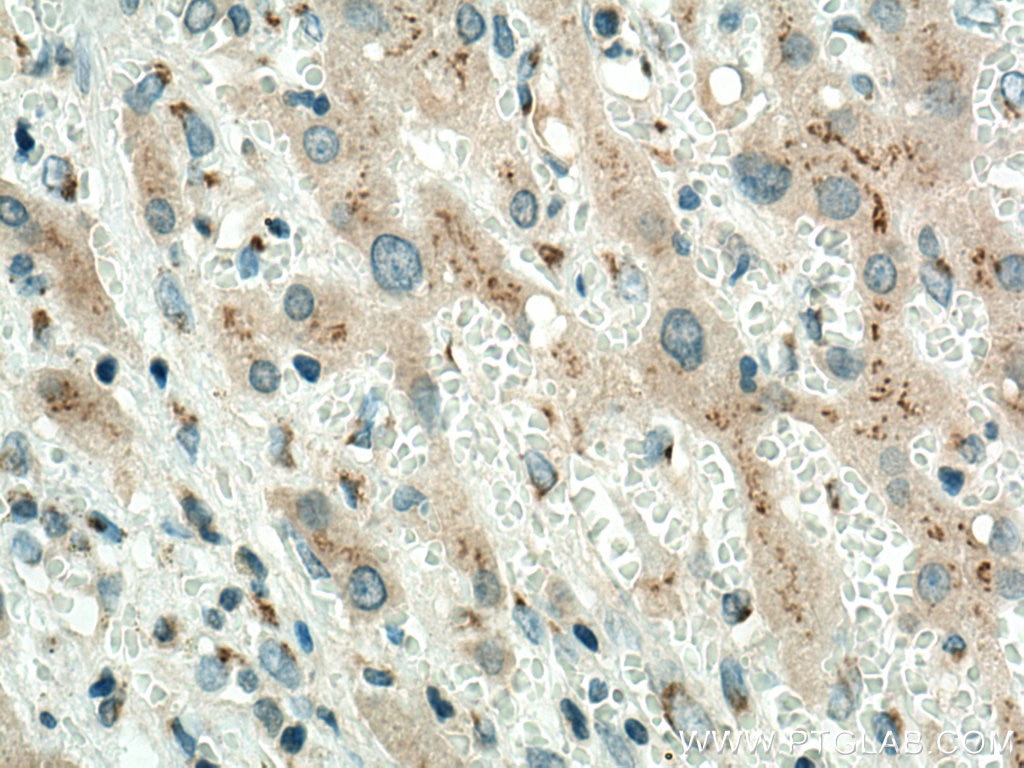
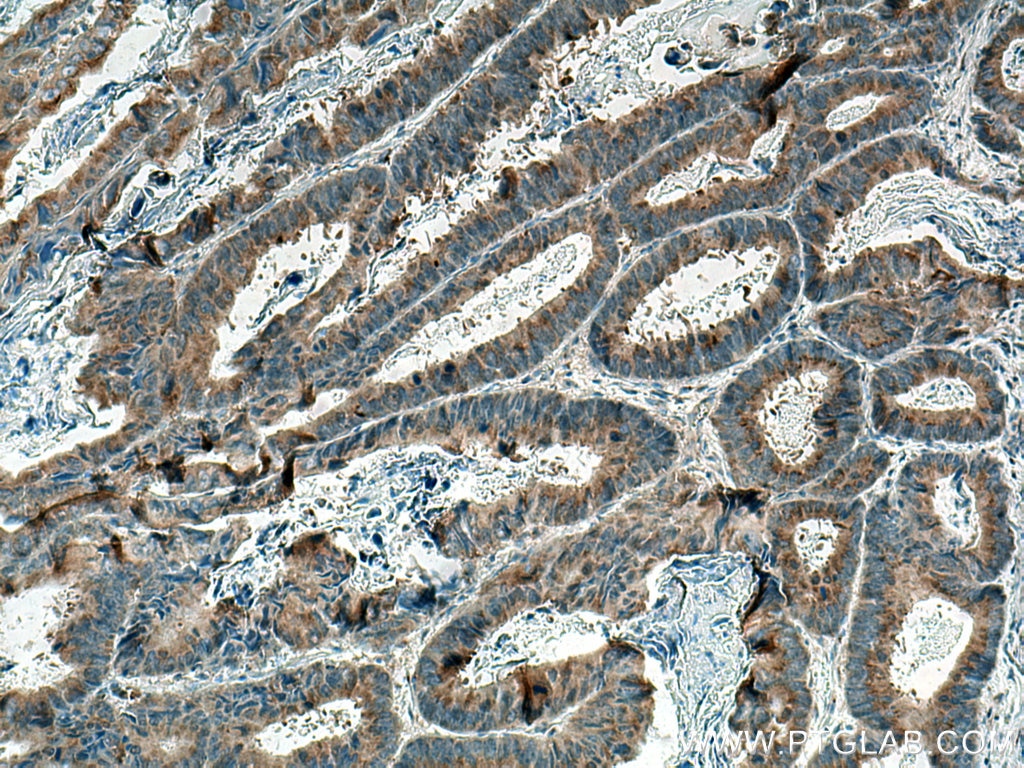
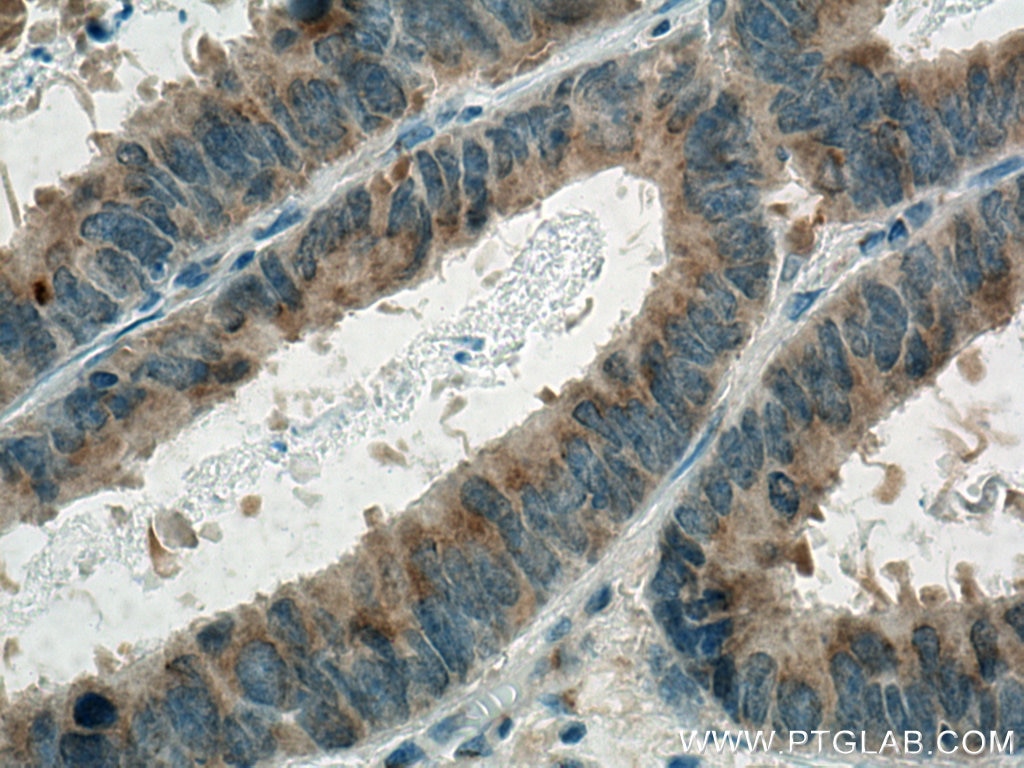
| Positive IHC detected in | human colon cancer tissue, human liver cancer tissue Note: suggested antigen retrieval with TE buffer pH 9.0; (*) Alternatively, antigen retrieval may be performed with citrate buffer pH 6.0 |
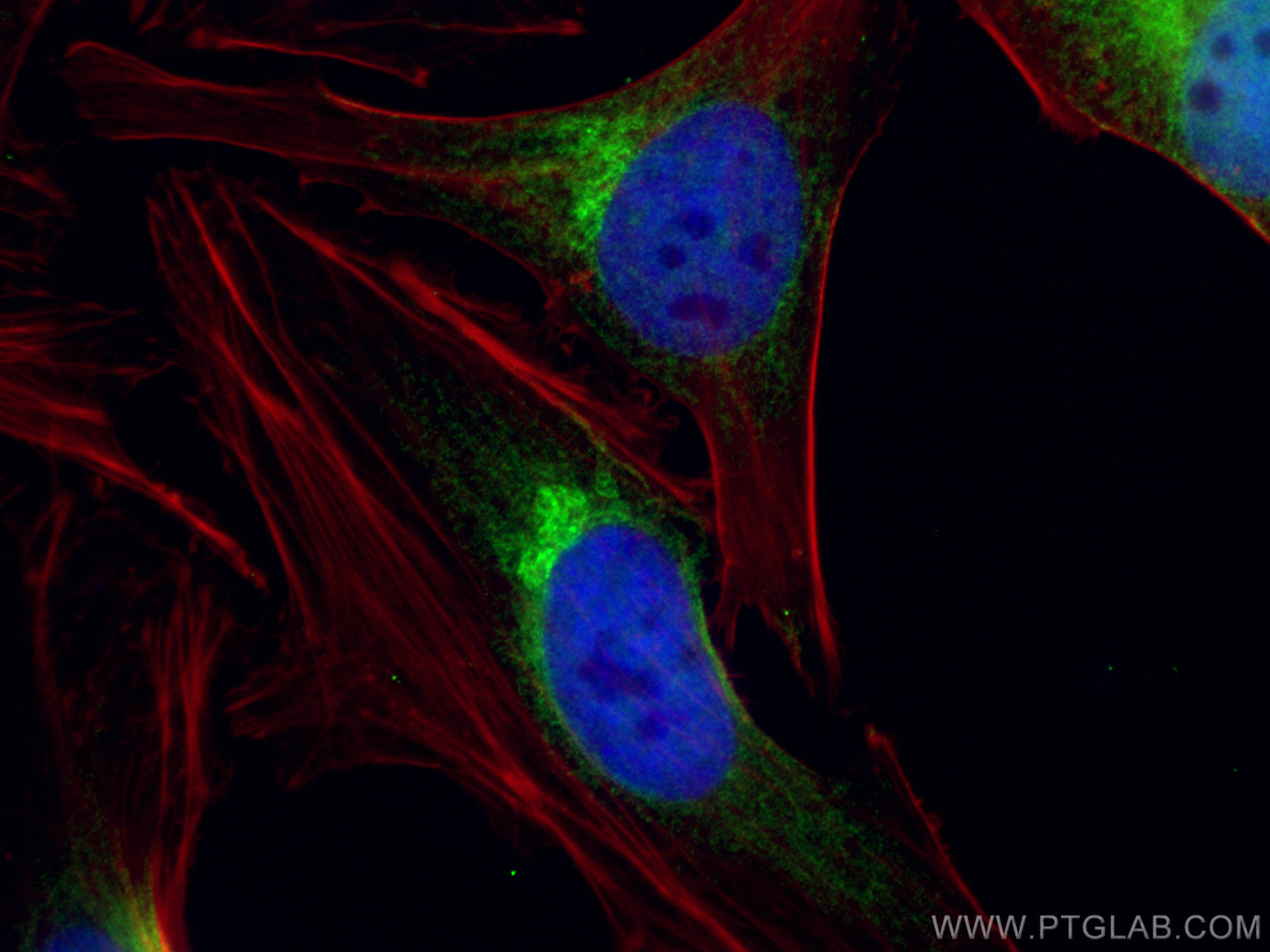
| Positive IF/ICC detected in | HeLa cells |
Recommended dilution
| Application | Dilution |
|---|---|
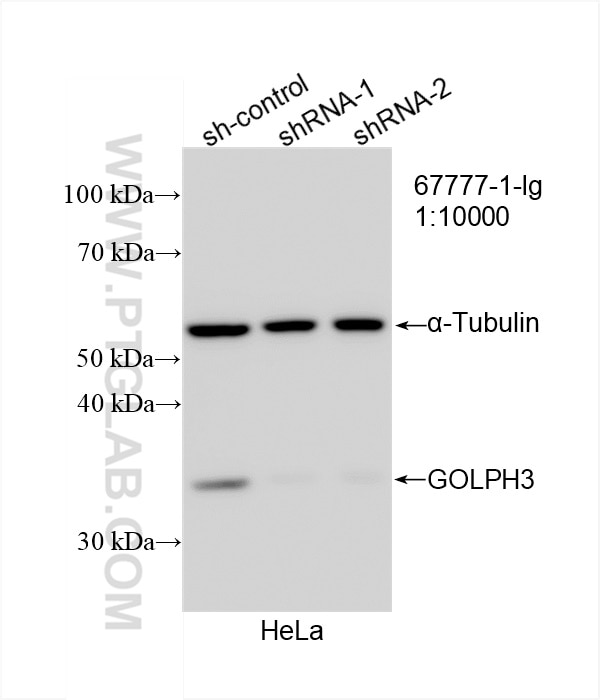
| Western Blot (WB) | WB : 1:5000-1:50000 |
| Immunohistochemistry (IHC) | IHC : 1:500-1:2000 |
| Immunofluorescence (IF)/ICC | IF/ICC : 1:200-1:800 |
| It is recommended that this reagent should be titrated in each testing system to obtain optimal results. | |
| Sample-dependent, Check data in validation data gallery. | |
Published Applications
| IF | See 2 publications below |
Product Information
67777-1-Ig targets GOLPH3 in WB, IHC, IF/ICC, ELISA, PLA applications and shows reactivity with human, mouse, rat samples.
| Tested Reactivity | human, mouse, rat |
| Cited Reactivity | human, mouse, rat |
| Host / Isotype | Mouse / IgG2a |
| Class | Monoclonal |
| Type | Antibody |
| Immunogen |
CatNo: Ag5443 Product name: Recombinant human GOLPH3 protein Source: e coli.-derived, PET28a Tag: 6*His Domain: 1-298 aa of BC033725 Sequence: MTSLTQRSSGLVQRRTEASRNAADKERAAGGGAGSSEDDAQSRRDEQDDDDKGDSKETRLTLMEEVLLLGLKDREGYTSFWNDCISSGLRGCMLIELALRGRLQLEACGMRRKSLLTRKVICKSDAPTGDVLLDEALKHVKETQPPETVQNWIELLSGETWNPLKLHYQLRNVRERLAKNLVEKGVLTTEKQNFLLFDMTTHPLTNNNIKQRLIKKVQEAVLDKWVNDPHRMDRRLLALIYLAHASDVLENAFAPLLDEQYDLATKRVRQLLDLDPEVECLKANTNEVLWAVVAAFTK Predict reactive species |
| Full Name | golgi phosphoprotein 3 (coat-protein) |
| Calculated Molecular Weight | 298 aa, 34 kDa |
| Observed Molecular Weight | 34 kDa |
| GenBank Accession Number | BC033725 |
| Gene Symbol | GOLPH3 |
| Gene ID (NCBI) | 64083 |
| RRID | AB_2918542 |
| Conjugate | Unconjugated |
| Form | Liquid |
| Purification Method | Protein A purification |
| UNIPROT ID | Q9H4A6 |
| Storage Buffer | PBS with 0.02% sodium azide and 50% glycerol, pH 7.3. |
| Storage Conditions | Store at -20°C. Stable for one year after shipment. Aliquoting is unnecessary for -20oC storage. 20ul sizes contain 0.1% BSA. |
Background Information
GOLPH3 (also called GPP34, GMx33, MIDAS, or yeast Vps74p) is a 34-kDa Golgi-associated protein conserved from yeast to human. GOLPH3 binds to PtdIns(4)P-rich trans-Golgi membranes and MYO18A conveying a tensile force required for efficient tubule and vesicle formation (PMID: 19837035). GOLPH3 has been recently demonstrated as a novel oncoprotein amplified in various types of human malignancies, including melanoma, breast, non-small cell lung cancer, gliomas and connective tissue tumors (PMID:19553991; 23006319; 21499727; 22745132). Enhanced activation of mTOR signaling represents a molecular basis for the oncogenic activity of GOLPH3 (PMID: 19553991).
Protocols
| Product Specific Protocols | |
|---|---|
| IF protocol for GOLPH3 antibody 67777-1-Ig | Download protocol |
| IHC protocol for GOLPH3 antibody 67777-1-Ig | Download protocol |
| WB protocol for GOLPH3 antibody 67777-1-Ig | Download protocol |
| Standard Protocols | |
|---|---|
| Click here to view our Standard Protocols |
Publications
| Species | Application | Title |
|---|---|---|
Cell Death Dis GOLPH3/CKAP4 promotes metastasis and tumorigenicity by enhancing the secretion of exosomal WNT3A in non-small-cell lung cancer | ||
J Adv Res Inflammation-driven biomimetic nano-polyphenol drug delivery system alleviates severe acute pancreatitis by inhibiting macrophage PANoptosis and pancreatic enzymes oversecretion | ||
Adv Sci (Weinh) Sequential Targeting Chondroitin Sulfate-Bilirubin Nanomedicine Attenuates Osteoarthritis via Reprogramming Lipid Metabolism in M1 Macrophages |
Reviews
The reviews below have been submitted by verified Proteintech customers who received an incentive for providing their feedback.
FH Boyan (Verified Customer) (12-06-2022) | No specific staining at all.
|