Autophagy in the Pathogenesis of Disease
Aging, cancer, inflammation, infection, the microenvironment, and neurodegenerative disease
| Autophagy Signaling Pathway | Neurodegenerative diseases |
| Human Diseases and Autophagy | Infectious and inflammatory diseases |
| Cancer and the microenvironment | References |
Download: Autophagy Research Focus PDF |
Autophagy Signaling Pathway
The 2016 Nobel Prize winner, Yoshinori Ohsumi, described autophagy as “a fundamental and an essential process for cellular health that plays a critical role in cell homeostasis, differentiation, nutrient deprivation, and normal growth control” (1).
The process of autophagy flux has five steps: autophagy induction, vesicle nucleation, expansion and completion, lysosome fusion (autophagolysosome formation), and breakdown (degradation & recycling). This mechanism is tightly regulated by mTOR, ULK1 complex, and ATG molecules, indicating significant crosstalk between signaling pathways ((2); Figure 1).
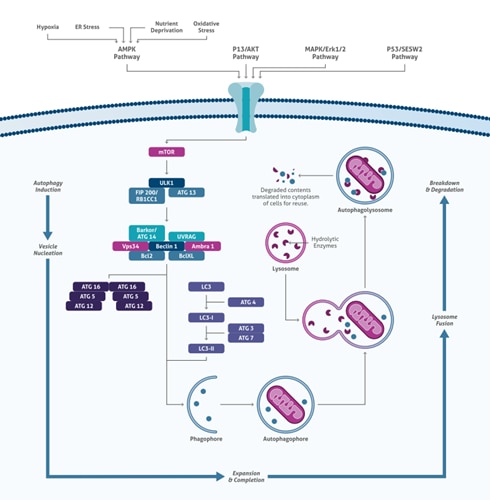
Figure 1. Multiple factors, including hypoxic and endoplasmic reticulum (ER) stress, nutrient deprivation, and oxidative stress, are involved in autophagy regulation.
Human Diseases and Autophagy
Aging process
The accumulation of damaged proteins is the main reason for age-associated loss of cellular function. The role of autophagy in the mobilization of lipid droplets, tumor development, and microbial pathogenesis has been extended to conditions related to the loss of autophagy during the aging process (3).
Autophagy is also implicated in different diseases, including neurodegenerative disorders, immune disease, and cancer ((4); Table 1).
| Genes | Functions related to autophagy | Human-associated diseases |
| ATG5 | Autophagosome formation | Genetic polymorphisms associated with asthma and systemic lupus erythematosus |
| ATG16L1 | Autophagosome formation | Mutations associated with Crohn’s disease |
| BECN1 | Autophagosome formation | Monoallelic deletion associated with human breast, ovarian, prostate, and colorectal cancers |
| EI24/PIG8 | Autophagosome formation and/or degradation | Mutations and deletions associated with the onset of human breast cancers |
| IRGM | Phagosome degradation | Single-nucleotide polymorphisms (SNPs) and deletion mutation associated with Crohn’s disease |
| NOD2/CARD15 | Autophagosome formation | SNPs and mutational variants associated with Crohn’s disease |
| PARK2/Parkin | Mitophagy induction / escape autophagic degradation | Mutations associated with autosomal recessive or sporadic Parkinson’s disease |
| PARK6/PINK1 | Mitophagy induction | Mutations associated with autosomal recessive or sporadic Parkinson’s disease |
| SMURF1 | Mediator of viral autophagy and mitophagy | SNP variants associated with ulcerative colitis |
| SQSTM1/p62 | An autophagy receptor for degradation of ubiquitinated substrates | Mutations associated with Paget’s disease and amyotrophic lateral sclerosis (ALS) |
| TECPR2 | Autophagosome formation | A frameshift mutation associated with hereditary spastic paraparesis |
| UVRAG | Autophagosome degradation | Deletion mutation associated with human colorectal cancer |
| WDR45/WIPI4 | Autophagosome formation | Heterozygous mutations associated with static encephalopathy of childhood with neurodegeneration in adulthood (SENDA) |
Table 1. Human diseases caused by defects in the autophagic machinery.
Autophagy, Cancer, and the microenvironment
The role of autophagy in cancer is complex and paradoxical. Beclin 1 (Atg 6; Figure 2) plays a crosstalk role between apoptosis autophagy and cancer. Beclin 1 is a mammalian tumor suppressor, detected monoallelically in approx 75% of ovarian, 50% of breast, and 40% of prostate cancers.
While autophagic deficiency has been shown to promote tumorigenesis in animal models, autophagy may support tumor growth by enhancing cancer cell survival in the face of nutrient depletion or accumulation of toxic molecules. It can also modulate the tumor microenvironment by promoting angiogenesis, nutrient supply, or modulating the inflammatory response. It implies that cancer stem cells (CSCs) may not be as rare as previously estimated, at the same time as highlighting the vital role of the microenvironment in supporting tumor growth.
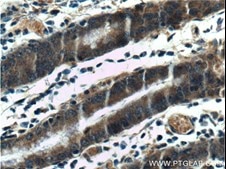 |
Figure 2. IHC of paraffin-embedded human stomach tissue slide using Beclin 1 Antibody (11306-1-AP); 1:200 and under a 40x lens. |
Neurodegenerative diseases and Autophagy
Since the autophagic pathway promotes cell survival by removing unwanted cellular organelle and protein aggregates, autophagy defects in neural cells can lead to neurodegeneration (5). These substrates reach lysosomes by several distinct mechanisms, including endosomal delivery or autophagosomes. Neurons are particularly vulnerable to disruptions of these interactions, especially as the brain ages. Unsurprisingly, mutations in genes regulating autophagy cause neurodegenerative diseases such as Alzheimer’s, amyotrophic lateral sclerosis (ALS), and familial Parkinson’s disease (Figure 3). Fortunately, the autophagy pathway has numerous druggable targets that may be able to combat these neurodegenerative diseases.
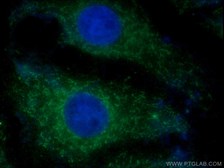 |
Figure 3. IF analysis of SH-SY5Y cells using Parkin Antibody (14060-1-AP); 1:50 and Alexa Fluor 488-congugated AffiniPure Goat Anti-Rabbit IgG(H+L). |
Autophagy and infectious/inflammatory diseases.
Autophagy also plays an important role in immune defense against bacteria and pathogens. During infection, autophagy regulates inflammation, antigen presentation, and micro-organism capture and degradation (6). Additionally, several immune mediators induce or repress autophagy. Moreover, immune signaling cascades are subject to regulation by autophagy, and a return to homeostasis following a robust immune response is critically dependent on this pathway. Improved mechanistic understanding of the autophagy machinery offers hope for treating infectious and inflammatory diseases.
Final remarks
When it was first discovered, autophagy was thought to be simply a garbage truck for cellular components. However, it is now recognized as a powerful tool in the cell, which can be used for good (protection of the cell) or ill (cancer and neurodegeneration). A better understanding of autophagy and autophagy modulators is important for the development of autophagy-based therapeutics that could treat various human diseases.
References:
1) Medicine Nobel for research on how cells 'eat themselves'.
2) Regulation Mechanisms and Signaling Pathways of Autophagy.
4) Autophagy and human diseases.
5) The role of autophagy in neurodegenerative disease.
6) Crosstalk between autophagy and inflammatory signaling pathways: balancing defense and homeostasis.
